Assessing the fitness of Epstein-Barr virus following its reactivation
- PMID: 40444990
- PMCID: PMC12282151
- DOI: 10.1128/jvi.00626-25
Assessing the fitness of Epstein-Barr virus following its reactivation
Abstract
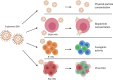
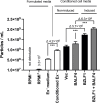
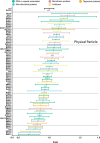
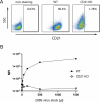
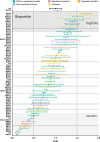
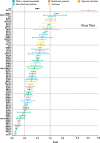
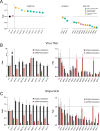
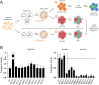
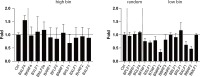
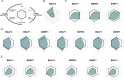
Epstein-Barr virus (EBV) is usually maintained latently on passaging in cell culture, as are most herpesviruses in vivo. Its fitness, or its ability to infect and replicate in naïve cells, cannot be ascertained by serially passaging it in host cells because no identified cell line initially supports a productive infection by it. Yet its fitness is critical to EBV's remarkable success as a human pathogen. We have, therefore, developed multiple approaches to assess EBV's fitness upon being reactivated from its familiar state of latency. We established and tested expression plasmids for 77 viral genes and a set of shRNAs targeting 25 viral genes to measure how increasing and decreasing their levels affected the fitness of the released stocks of virus. Four of their properties were then analyzed: (i) their concentrations of physical particles, (ii) their binding to the CD21 receptor on a B-cell line, (iii) their entry into human primary B cells, and (iv) their infectious titers. These analyses identified multiple EBV genes whose altered levels of expression altered the biological activities of the released virus. These measurements revealed, though, an unexpected insight into the robustness of EBV produced from latently infected cells. EBV is amazingly resilient to any increased expression of its genes. The levels expressed in cells, as they support an induced productive infection, therefore are close to optimal for the fitness of the released virus.IMPORTANCEPopulations of viruses accumulate mutations while being propagated. While most mutations are neutral or disadvantageous, some confer on the variant a selective advantage, increasing its infectivity. These variants can be identified by serial passaging virus stocks, allowing those with increased fitness to predominate. This approach does not work for Epstein-Barr virus (EBV), for which no identified cell line initially supports its productive infection. How mutations accumulate in EBV as it is propagated latently to affect its fitness was unknown. We have devised an approach to assess EBV's fitness upon being reactivated. Our findings suggest that EBV during its many latent generations has maintained a strikingly robust productive fitness.
Keywords: EBV; Epstein-Barr virus; fusion assay; herpesvirus; infection; mutagenesis; physical particles; viral fitness; virus titer.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

