Striatal cell-type-specific molecular signatures reveal potential therapeutic targets in a model of dystonia
- PMID: 40447200
- PMCID: PMC12223790
- DOI: 10.1016/j.nbd.2025.106981
Striatal cell-type-specific molecular signatures reveal potential therapeutic targets in a model of dystonia
Abstract
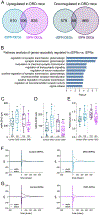
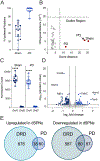
Abnormal dopamine neurotransmission and striatal dysfunction is implicated in many forms of dystonia, yet the underlying molecular processes remain unknown. Here, we identified thousands of dysregulated genes within striatal spiny projection neuron (SPN) subtypes in a genetic mouse model of DOPA-responsive dystonia (DRD), which is caused by gene defects that reduce dopamine neurotransmission. Although changes in mRNA expression were unique to each SPN subtype, abnormal glutamatergic signaling was implicated in each SPN subtype. Indeed, both AMPA and NMDA receptor-mediated currents were enhanced in direct SPNs but diminished in indirect SPNs in DRD mice. The pattern of mRNA dysregulation was distinct from parkinsonism where the dopamine deficit occurs in adults, suggesting that the phenotypic outcome is dependent on both the timing of the dopaminergic deficit and the SPN-specific adaptions. By leveraging these disease-specific molecular signatures, we identified LRRK2 inhibition, among other mechanisms, as a novel therapeutic target for dystonia.
Keywords: D1 dopamine receptor; D2 dopamine receptor; MLi-2; Medium spiny neuron; Parkinson's disease; RNA-seq; Translatome.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare the following financial interests/personal relationships which may be considered as potential competing interests:(Ellen J Hess reports financial support was provided by National Institutes of Health. C. Savio Chan reports financial support was provided by National Institutes of Health. Ellen J Hess reports financial support was provided by Parkinson's Foundation Inc. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.)
Figures



Update of
-
Striatal cell-type-specific molecular signatures reveal therapeutic targets in a model of dystonia.bioRxiv [Preprint]. 2024 Oct 7:2024.10.07.617010. doi: 10.1101/2024.10.07.617010. bioRxiv. 2024. Update in: Neurobiol Dis. 2025 Aug;212:106981. doi: 10.1016/j.nbd.2025.106981. PMID: 39415987 Free PMC article. Updated. Preprint.
References
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

