Genome-wide analysis of the Tritipyrum bHLH gene family and the response of TtbHLH310 in salt-tolerance
- PMID: 40448022
- PMCID: PMC12123985
- DOI: 10.1186/s12864-025-11657-z
Genome-wide analysis of the Tritipyrum bHLH gene family and the response of TtbHLH310 in salt-tolerance
Abstract
Background: The bHLH transcription factor is prevalent across the plant kingdom and is crucial for various abiotic stress responses in different plant species. Tritipyrum, an octoploid created from an intergeneric cross between Triticum aestivum (AABBDD) and Thinopyrum elongatum (EE), serves as a significant source of germplasm, facilitating the incorporation of desirable traits from Th. elongatum into T. aestivum. With the recent availability of the complete genome sequences of T. aestivum and Th. elongatum, it has become feasible to investigate the organization and expression patterns of bHLH genes within the Tritipyrum genome.
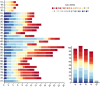
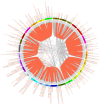
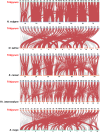
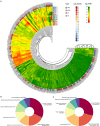
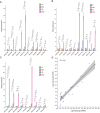
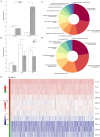
Results: In this study, a total of 398 bHLH genes (TtbHLH) were identified within the Tritipyrum genome. These genes were classified into twenty major groups based on evolutionary analysis, indicating that they share conserved motif compositions. The TtbHLH genes are distributed across 28 chromosomes and include 67 duplication events. Synteny analysis suggests a common ancestral lineage for the bHLH gene family. Transcriptome data and quantitative polymerase chain reaction (qPCR) expression profiling identified 29 TtbHLH genes with significantly elevated expression levels in response to various salt-stress conditions and recovery treatments. Notably, Tel1E01T336100 (TtbHLH310) demonstrated a pronounced sensitivity to salt stress and is phylogenetically related to the salt-tolerant gene AtbHLH6 in Arabidopsis thaliana. Additionally, Pearson correlation analysis revealed 485 genes that exhibited a strong positive correlation (R > 0.9) with TtbHLH310 expression, which was enriched in pathways related to metabolic activities, cellular processes, stimulus responses, and biological regulation. Further analysis through real-time PCR confirmed that TtbHLH310 is highly expressed in the roots, stems, and leaves under salt-stress conditions.
Conclusions: The findings indicate that TtbHLH310 may play a pivotal role in enhancing salt stress tolerance in plants. Its strong expression in response to salt stress highlights its potential as a valuable foreign gene for improving salt tolerance in wheat. These insights contribute to our understanding of the molecular mechanisms underpinning abiotic stress responses in Tritipyrum and may aid in the development of more resilient wheat varieties.
Keywords: Tritipyrum; TtbHLH310; Expression patterns; Genome-wide; Salt-tolerance; bHLH.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: This article does not contain any studies with human participants or animals performed by the authors. These methods were carried out in accordance with relevant guidelines and regulations. We confirm that all experimental protocols were approved by Anshun University. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures
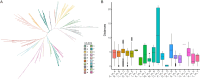
References
-
- Li J, Pu L, Han M, Zhu M, Zhang R, Xiang Y. Soil salinization research in China: advances and prospects. J Geog Sci. 2014;24(5):943–60. - DOI
-
- Qadir M, Quillérou E, Nangia V, Murtaza G, Singh M, Thomas RJ, Drechsel P, Noble AD. Economics of salt-induced land degradation and restoration. Nat. Resour. Forum. 2014;38(4): 282–95.
MeSH terms
Substances
Grants and funding
- No. 32460493/The Regional Science Fund Project of the National Natural Science Foundation of China
- QKFQ[2022]007/The Innovation Capacity Building Project of Guizhou Scientific Institutions
- QNKZZZY(2023)06/Guizhou Academy of Agricultural Sciences Key Laboratory Project for Crop Gene Resources and Germplasm Innovation in Karst Mountain regions
- Qiankehe Basic-ZK [2023] Key No.002/The Science and Technology Department of Guizhou
LinkOut - more resources
Full Text Sources

