Early transcriptional and cellular abnormalities in choroid plexus of a mouse model of Alzheimer's disease
- PMID: 40450296
- PMCID: PMC12125878
- DOI: 10.1186/s13024-025-00853-w
Early transcriptional and cellular abnormalities in choroid plexus of a mouse model of Alzheimer's disease
Abstract
Background: Alzheimer's disease (AD) is a progressive neurodegenerative disorder characterized by the accumulation of amyloid-β plaques, tau hyperphosphorylation, and neuroinflammation. The choroid plexus (ChP), serving as the blood-cerebrospinal fluid-brain barrier, plays essential roles in immune response to stress and brain homeostasis. However, the cellular and molecular contributions of the ChP to AD progression remain inadequately understood.
Methods: To elucidate the molecular abnormalities during the early stages of AD, we acquired single-cell transcription profiling of ChP from APP/PS1 mice with early-stage of Aβ pathology and litter-mate controls. The transcriptional alterations that occurred in each cell type were identified by differentially expressed genes, cell-cell communications and pseudotemporal trajectory analysis. The findings were subsequently validated by a series of in situ and in vitro assays.
Results: We constructed a comprehensive atlas of ChP at single-cell resolution and identified six major cell types and immune subclusters in male mice. The majority of dysregulated genes were found in the epithelial cells of APP/PS1 mice in comparison to wild-type (WT) mice, and most of these genes belonged to down-regulated module involved in mitochondrial respirasome assembly, cilium organization, and barrier integrity. The disruption of the epithelial barrier resulted in the downregulation of macrophage migration inhibitory factor (MIF) secretion in APP/PS1 mice, leading to macrophage activation and increased phagocytosis of Aβ. Concurrently, ligands (e.g., APOE) secreted by macrophages and other ChP cells facilitated the entry of lipids into ependymal cells, leading to lipid accumulation and the activation of microglia in the brain parenchyma in APP/PS1 mice compared to WT controls.
Conclusions: Taken together, these data profiled early transcriptional and cellular abnormalities of ChP within an AD mouse model, providing novel insights of cerebral vasculature into the pathobiology of AD.
Keywords: Alzheimer’s disease; Choroid plexus; Lipid accumulation; Neuroinflammation; Single-cell transcriptome.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All animal experiments in this study were approved by the Kunming Institute of Zoology and strictly adhered to the animal care regulations of the Institutional Animal Care and Use Committee of Kunming Institute of Zoology. Consent for publication: All authors have read the final draft of the manuscript and approved its submission to Molecular Neurodegeneration. Competing interests: We declare that there is no conflict of interest and the manuscript has not been published, accepted for publication, or placed under any editorial review for publication elsewhere.
Figures


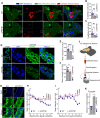


References
MeSH terms
Substances
Grants and funding
- STI2030-Major Projects-2022ZD0213500/Ministry of Science and Technology of the People's Republic of China
- 2021ZD0200901/Ministry of Science and Technology of the People's Republic of China
- 32230021/National Natural Science Foundation of China
- 82022017/National Natural Science Foundation of China
- 202305AH340006/Applied Basic Research Foundation of Yunnan Province
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

