Inhibition of acid or neutral sphingomyelinases differentially impacts RNA and protein cargo sorting to extracellular vesicles
- PMID: 40454089
- PMCID: PMC12124672
- DOI: 10.1016/j.isci.2025.112440
Inhibition of acid or neutral sphingomyelinases differentially impacts RNA and protein cargo sorting to extracellular vesicles
Abstract
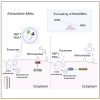
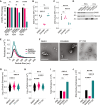
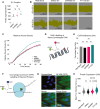
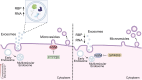
Extracellular vesicles (EVs) form through regulated biogenesis processes involving sphingomyelinases (SMases), enzymes that metabolize sphingomyelin to produce ceramide-a lipid influencing membrane rigidity and essential for EV generation. This study explores the impact of inhibiting neutral SMase (NSM) and acid SMase (ASM) on the sorting of EV protein and RNA cargoes in human MCF7 cells. Our results revealed that NSM inhibition reduces EV nanoparticles and diminishes RNA and protein cargoes, including endosomal, spliceosomal, and translation-related proteins. Conversely, ASM inhibition increased RNA-binding proteins within and enhanced the expression of ribonucleoprotein complex-associated RNA in released EVs, including several snRNAs and 7SL RNA. Intriguingly, ASM-inhibited EVs enhanced the migration and translational activity of recipient MCF10A cells. These findings suggest an important role for SMase-dependent vesiculation in governing RNA and protein trafficking to the extracellular space, unveiling potential implications for cellular communication and function.
Keywords: Biological sciences; Cell biology; Omics; Organizational aspects of cell biology.
© 2025 The Authors. Published by Elsevier Inc.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
LinkOut - more resources
Full Text Sources

