Comparative repeatome analysis of Pyrgomorphidae and Acrididae (Orthoptera: Caelifera) revealed the contribution of repetitive DNA in genome gigantism
- PMID: 40455812
- PMCID: PMC12129174
- DOI: 10.1371/journal.pone.0325165
Comparative repeatome analysis of Pyrgomorphidae and Acrididae (Orthoptera: Caelifera) revealed the contribution of repetitive DNA in genome gigantism
Abstract
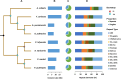
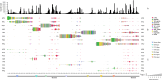
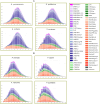
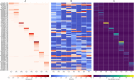
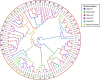
Eukaryotic genomes are often rich in DNA repetitive elements, involving both transposable elements (TE) and tandemly repeated satellite DNA. Grasshopper species, known for their large genome sizes, comprising relatively a high proportion of genomic repeats. This study aimed to identify and perform a comparative analysis of DNA repetitive content in eight grasshopper species from the Pyrgomorphidae and Acrididae families. We utilized unassembled low-coverage Illumina paired-end short reads in the RepeatExplorer2 pipeline to identify genomic repeats, and RepeatMasker to estimate their abundance and divergence activity. Flow cytometry estimated genome sizes, ranging from 1C = 7.670 pg to 18.612 pg, with Aularches miliaris (18.612 pg) being the second largest insect genome reported to date. The repeat content ranged from 51% to 74%, with a mean value of 64.26% of the total genome. The major identified repeat elements included LINE, Ty3_Gypsy, Penelope, Ty1-copia, Helitron, Maverick, and satellite repeats, with LINE elements being the most abundant, constituting 24% to 54% of the total repetitive content in Apalacris varicornis and A. miliaris, respectively. The positive correlation of repetitive content and TEs with genome size suggests that their expansion has contributed to the large genome sizes observed. Satellite DNA analysis identified 65 satDNA families across the eight species. Additionally, phylogenetic analysis of TE protein domains revealed that consensus sequences from the same domain cluster together, suggesting domain-specific evolutionary pathways for TEs in the genome. This study reveals new dynamics into the role of repetitive DNA in genome gigantism as well as other evolutionary mechanisms in the Pyrgomorphidae and Acrididae families of Orthoptera.
Copyright: © 2025 Salman et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


Similar articles
-
Comparative analysis of morabine grasshopper genomes reveals highly abundant transposable elements and rapidly proliferating satellite DNA repeats.BMC Biol. 2020 Dec 21;18(1):199. doi: 10.1186/s12915-020-00925-x. BMC Biol. 2020. PMID: 33349252 Free PMC article.
-
Comparative Analysis of Genomic Repeat Content in Gomphocerine Grasshoppers Reveals Expansion of Satellite DNA and Helitrons in Species with Unusually Large Genomes.Genome Biol Evol. 2020 Jul 1;12(7):1180-1193. doi: 10.1093/gbe/evaa119. Genome Biol Evol. 2020. PMID: 32539114 Free PMC article.
-
Repetitive element expansions contribute to genome size gigantism in Pamphagidae: A comparative study (Orthoptera, Acridoidea).Genomics. 2024 Sep;116(5):110896. doi: 10.1016/j.ygeno.2024.110896. Epub 2024 Jul 19. Genomics. 2024. PMID: 39025318
-
The repetitive DNA content of eukaryotic genomes.Genome Dyn. 2012;7:1-28. doi: 10.1159/000337118. Epub 2012 Jun 25. Genome Dyn. 2012. PMID: 22759811 Review.
-
Structural and functional liaisons between transposable elements and satellite DNAs.Chromosome Res. 2015 Sep;23(3):583-96. doi: 10.1007/s10577-015-9483-7. Chromosome Res. 2015. PMID: 26293606 Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

