Deciphering genetic adaptations of Old World camels through comparative genomic analyses across all camelid species
- PMID: 40458189
- PMCID: PMC12127551
- DOI: 10.1016/j.isci.2025.112477
Deciphering genetic adaptations of Old World camels through comparative genomic analyses across all camelid species
Abstract
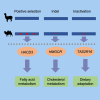
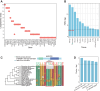
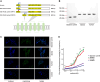
Old World camels exhibit many unique traits for adaptations to desert environments, distinct from New World camels within the same family. Here, we conducted a comparative genomic analysis of three Old World camel species relative to four New World camel species, as well as cattle, pigs, mice, and humans to search for genes showing positive selection and specific variations in Old World camels. We identified genes under positive selection, genes with specific small indels, and inactivated genes associated with the unique lipid metabolism and skin characteristics of Old World camels. Especially, through experimental validation, we confirmed the inactivation of TAS2R16, a gene encoding a prominent bitter taste receptor, which could enable Old World camels to consume certain toxic plants with a bitter taste to other mammals. These findings provide insights into the genetic basis underlying the distinctive energy metabolism, water-salt homeostasis, and dietary adaptations of Old World camels.
Keywords: Evolutionary biology; Evolutionary ecology; Zoology.
© 2025 The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Mitochondrial DNA variation and phylogeography of Old World camels.Anim Biosci. 2021 Apr;34(4):525-532. doi: 10.5713/ajas.20.0319. Epub 2020 Aug 24. Anim Biosci. 2021. PMID: 32898955 Free PMC article.
-
Signatures of purifying selection and site-specific positive selection on the mitochondrial DNA of dromedary camels (Camelus dromedarius).Mitochondrion. 2023 Mar;69:36-42. doi: 10.1016/j.mito.2023.01.004. Epub 2023 Jan 20. Mitochondrion. 2023. PMID: 36690316
-
Development and validation of the Axiom-MaruPri SNP chip for genetic analyses of domesticated old world camelids.Gene. 2024 Aug 30;921:148541. doi: 10.1016/j.gene.2024.148541. Epub 2024 May 8. Gene. 2024. PMID: 38723784
-
Old World camels in a modern world - a balancing act between conservation and genetic improvement.Anim Genet. 2019 Dec;50(6):598-612. doi: 10.1111/age.12858. Epub 2019 Sep 18. Anim Genet. 2019. PMID: 31532019 Free PMC article. Review.
-
Review on camel genetic diversity: ecological and economic perspectives.Mamm Genome. 2024 Dec;35(4):621-632. doi: 10.1007/s00335-024-10054-3. Epub 2024 Jul 29. Mamm Genome. 2024. PMID: 39075281 Review.
References
-
- Balmus G., Trifonov V.A., Biltueva L.S., O'Brien P.C.M., Alkalaeva E.S., Fu B., Skidmore J.A., Allen T., Graphodatsky A.S., Yang F., Ferguson-Smith M.A. Cross-species chromosome painting among camel, cattle, pig and human: further insights into the putative Cetartiodactyla ancestral karyotype. Chromosome Res. 2007;15:499–515. doi: 10.1007/s10577-007-1154-x. - DOI - PubMed
-
- Mirgani T. Effect of Fasting on Camel Tissue Lipid. J. Arid Environ. 1981;4:359–361. doi: 10.1016/S0140-1963(18)31480-0. - DOI
-
- Guo F., Si R., He J., Yuan L., Hai L., Ming L., Yi L., Ji R. Comprehensive transcriptome analysis of adipose tissue in the Bactrian camel reveals fore hump has more specific physiological functions in immune and endocrine systems. Livest. Sci. 2019;228:195–200. doi: 10.1016/j.livsci.2019.09.003. - DOI
Associated data
LinkOut - more resources
Full Text Sources

