Therapeutic Targets for Sepsis: Multicenter Proteome-Wide Analyses and Experimental Validation
- PMID: 40459852
- PMCID: PMC12235691
- DOI: 10.1021/acs.jproteome.5c00148
Therapeutic Targets for Sepsis: Multicenter Proteome-Wide Analyses and Experimental Validation
Abstract
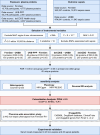
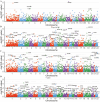
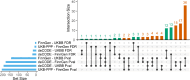
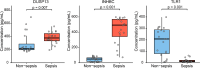
Sepsis is a life-threatening condition with high mortality, underscoring the urgent need for effective therapeutic targets. We conducted a proteome-wide Mendelian randomization (MR) analysis using plasma protein data from the FinnGen, UKB-PPP, and deCODE cohorts to identify proteins causally associated with sepsis. The analysis included 16,074 cases and 363,227 controls in FinnGen and 11,643 cases and 474,841 controls in the UK Biobank, spanning four exposure-outcome combinations. Proteins were prioritized based on a false discovery rate <0.05 in one combination and p < 0.05 in another. Colocalization and phenome-wide association studies (PheWAS) were performed to evaluate causality and potential off-target effects. Three proteins─dual specificity phosphatase 13 (DUSP13), inhibin beta C chain (INHBC), and toll-like receptor 1 (TLR1)─showed strong evidence of colocalization with sepsis risk. PheWAS confirmed broader disease associations for DUSP13 and TLR1, while INHBC showed no significant adverse associations and is considered druggable. TLR1 is currently under clinical investigation. ELISA-based experimental validation in 20 sepsis patients and 20 controls demonstrated elevated serum levels of DUSP13 and INHBC and reduced levels of TLR1 in sepsis. These findings identify DUSP13, INHBC, and TLR1 as promising therapeutic targets for sepsis, supported by genetic, phenotypic, and experimental evidence.
Keywords: PheWAS; colocalization analysis; druggability; proteome-wide Mendelian randomization; sepsis; therapeutic targets.
Figures

References
-
- World Health Organization. Sepsis, 2023. https://www.who.int/news-room/fact-sheets/detail/sepsis.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

