H3.3 deposition counteracts the replication-dependent enrichment of H3.1 at chromocenters in embryonic stem cells
- PMID: 40461574
- PMCID: PMC12134209
- DOI: 10.1038/s41467-025-60430-z
H3.3 deposition counteracts the replication-dependent enrichment of H3.1 at chromocenters in embryonic stem cells
Abstract
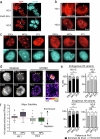
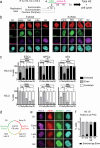
Chromocenters in mouse cells are membrane-less nuclear compartments representing typical heterochromatin stably maintained during cell cycle. We explore how histone H3 variants, replicative H3.1/2 or replacement H3.3, mark these domains during the cell cycle in mouse embryonic stem cells, neuronal precursor cells as well as immortalized 3T3 cells. We find a strong and distinct H3.1 enrichment at chromocenters, with variation in mouse embryonic stem cells. Mechanistically, this H3.1 selective enrichment depends on the DNA Synthesis Coupled deposition pathway operating in S phase challenged when we target H3.3 deposition through the DNA Synthesis Independent deposition pathway mediated by HIRA. Altering the H3.1/H3.3 dynamics at chromocenters in mouse embryonic stem cells affects nuclear morphology and cell division. Here, we reveal opposing mechanisms for H3.1 and H3.3 deposition with different enforcement according to cell cycle and potency which determine their ratio at chromocenters and are critical for genome stability and cell survival.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

