Acquired resistance in cancer: towards targeted therapeutic strategies
- PMID: 40461793
- PMCID: PMC12307123
- DOI: 10.1038/s41568-025-00824-9
Acquired resistance in cancer: towards targeted therapeutic strategies
Abstract
Development of acquired therapeutic resistance limits the efficacy of cancer treatments and accounts for therapeutic failure in most patients. How resistance arises, varies across cancer types and differs depending on therapeutic modalities is incompletely understood. Novel strategies that address and overcome the various and complex resistance mechanisms necessitate a deep understanding of the underlying dynamics. We are at a crucial time when innovative technologies applied to patient-relevant tumour models have the potential to bridge the gap between fundamental research into mechanisms and timing of acquired resistance and clinical applications that translate these findings into actionable strategies to extend therapy efficacy. Unprecedented spatial and time-resolved high-throughput platforms generate vast amounts of data, from which increasingly complex information can be extracted and analysed through artificial intelligence and machine learning-based approaches. This Roadmap outlines key mechanisms that underlie the acquisition of therapeutic resistance in cancer and explores diverse modelling strategies. Clinically relevant, tractable models of disease and biomarker-driven precision approaches are poised to transform the landscape of acquired therapy resistance in cancer and its clinical management. Here, we propose an integrated strategy that leverages next-generation technologies to dissect the complexities of therapy resistance, shifting the paradigm from reactive management to predictive and proactive prevention.
© 2025. Springer Nature Limited.
Conflict of interest statement
Competing interests: The authors declare no direct competing interests. For full disclosure, C.D.W. has received research funding from Varian Medical Systems, AACR-Novocure and OMS Foundation, clinical trial support from MuReva and Tactile Medical, and consultancy/honoraria from LifeNet Health, Guidepoint Global and EMD Serono; A.S. sits on the Board of the Society for Functional Precision Medicine; P.C.B. sits on the Scientific Advisory Boards of Intersect Diagnostics Inc., BioSymetrics Inc. and previously sat on the board of Sage Bionetworks; E.S.K. has sponsored research funded by Blueprint Medicines and Bristol Myers Squibb and is a member of the Cancer Cell Cyclse–LLC consulting enterprise; A.K.W. has sponsored research funded by Blueprint Medicines and Bristol Myers Squibb; C.E.T. has received funding from AstraZeneca; J.W.T. has received research support from Acerta, Agios, Aptose, Array, AstraZeneca, Constellation, Genentech, Gilead, Incyte, Janssen, Kronos, Meryx, Petra, Schrodinger, Seattle Genetics, Syros, Takeda and Tolero and serves on the advisory board for Recludix Pharm, AmMax Bio and Ellipses Pharma; V.C.S. is a consultant for and equity holder in Femtovox Inc.
Figures


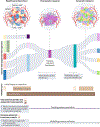
References
Publication types
MeSH terms
Substances
Grants and funding
- U24 CA274159/CA/NCI NIH HHS/United States
- U54 CA274220/CA/NCI NIH HHS/United States
- U01 CA223976/CA/NCI NIH HHS/United States
- P30 CA016042/CA/NCI NIH HHS/United States
- R01 CA244729/CA/NCI NIH HHS/United States
- R01 CA234162/CA/NCI NIH HHS/United States
- R01 CA256911/CA/NCI NIH HHS/United States
- R01 CA163649/CA/NCI NIH HHS/United States
- R37 CA276924/CA/NCI NIH HHS/United States
- R01 CA270234/CA/NCI NIH HHS/United States
- R01 CA264248/CA/NCI NIH HHS/United States
- R01 CA175495/CA/NCI NIH HHS/United States
- R01 CA175397/CA/NCI NIH HHS/United States
- R01 CA247362/CA/NCI NIH HHS/United States
- U01 CA271412/CA/NCI NIH HHS/United States
- U54 CA224019/CA/NCI NIH HHS/United States
- U54 CA224081/CA/NCI NIH HHS/United States
- U54 CA274329/CA/NCI NIH HHS/United States
- R01 CA240718/CA/NCI NIH HHS/United States
- R01 CA280980/CA/NCI NIH HHS/United States
- U24 CA248265/CA/NCI NIH HHS/United States
- U54 CA274321/CA/NCI NIH HHS/United States
- R01 CA270108/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

