Maternal loss of mouse Nlrp2 alters the transcriptome and DNA methylome in GV oocytes and impairs zygotic genome activation in embryos
- PMID: 40462191
- PMCID: PMC12135322
- DOI: 10.1186/s13148-025-01889-x
Maternal loss of mouse Nlrp2 alters the transcriptome and DNA methylome in GV oocytes and impairs zygotic genome activation in embryos
Abstract
Background: NLRP2 is a subcortical maternal complex (SCMC) protein of mammalian oocytes and preimplantation embryos. SCMC proteins are encoded by maternal effect genes and play a pivotal role in the maternal-to-zygotic transition (MZT), early embryogenesis, and epigenetic (re)programming. Maternal inactivation of genes encoding SCMC proteins has been linked to infertility and subfertility in mice and humans, but the underlying molecular mechanisms for the diverse functions of SCMC proteins, and specifically the role of NLRP2, are incompletely understood.
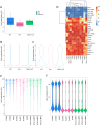
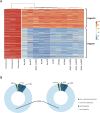
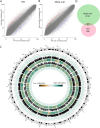
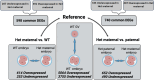
Results: We profiled the DNA methylome of pre-ovulatory germinal-vesicle (GV) oocytes from Nlrp2-null, heterozygous (Het), and wild-type (WT) female mice and assessed the transcriptome of GV oocytes and 2-cell embryos from WT and Nlrp2-null females. The absence or reduction of NLRP2 did not alter the distinctive global DNA methylation landscape of GV oocytes, including their unique bimodal methylome patterns and methylation at the germline differentially methylated regions (gDMRs) of imprinted genes. However, altered methylation was observed in a small subset of oocyte-characteristic hyper- and hypomethylated domains and within a minor fraction of genomic regions, particularly in Nlrp2-null oocytes. Transcriptome profiling revealed substantial differences between the Nlrp2-null and WT GV oocytes, including deregulation of many crucial factors involved in oocyte transcriptome modulation and epigenetic reprogramming. Moreover, maternal absence of NLRP2 significantly altered the transcriptome of heterozygous embryos from Nlrp2-null females compared to WT embryos, whereas the transcriptome of heterozygous embryos from Nlrp2-null males was not significantly different from that of WT embryos. Maternal absence of NLRP2 also negatively impacted MZT, as evidenced by the deregulation of a large subset of zygotic genome activation (ZGA)-related genes.
Conclusions: This study demonstrates that NLRP2 is essential for shaping the transcriptome of GV oocytes and preimplantation embryos. Maternal loss of Nlrp2 negatively impacts ZGA. Our findings that the DNA methylome of Het and Nlrp2-null oocytes was subtly changed, and that gene-body DNA methylation differences did not correlate with gene expression differences, suggest that posttranscriptional changes in transcript stability, rather than altered transcription itself, are primarily responsible for the changed transcriptome of Nlrp2-null oocytes.
Keywords: DNA methylation; Imprinted genes; Oocyte; Subcortical maternal complex.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: There is no human participant in this study. All experiments were approved by the Institutional Animal Care and Use Committee (IACUC) at Baylor College of Medicine (BCM) under study protocol (AN-2035) and conducted according to institutional and governmental regulations concerning the ethical use of animals. Consent for publication: Not applicable. Competing interests: The authors declare that they have no competing interests.
Figures




References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

