Multi-omics profiling of cross-resistance between ceftazidime-avibactam and meropenem identifies common and strain-specific mechanisms in Pseudomonas aeruginosa clinical isolates
- PMID: 40464559
- PMCID: PMC12239554
- DOI: 10.1128/mbio.03896-24
Multi-omics profiling of cross-resistance between ceftazidime-avibactam and meropenem identifies common and strain-specific mechanisms in Pseudomonas aeruginosa clinical isolates
Abstract
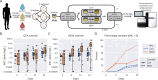
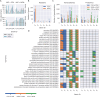
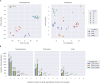
Pseudomonas aeruginosa is a highly versatile and resilient pathogen that can infect different tissues and rapidly develop resistance to multiple drugs. Ceftazidime-avibactam (CZA) is an antibiotic often used to treat multidrug-resistant infections; however, the knowledge on the CZA resistance mechanisms in P. aeruginosa is limited. Here, we performed laboratory evolution of eight clinical isolates of P. aeruginosa exposed to either CZA or meropenem (MEM) in sub-inhibitory concentrations and used multi-omics profiling to investigate emerging resistance mechanisms. The majority of strains exposed to MEM developed high resistance (83%, 20/24 strains from eight clinical isolates), with only 17% (4/24) acquiring cross-resistance to CZA. The rate of resistance evolution to CZA was substantially lower (21%, 5/24), while 38% (9/24) acquired cross-resistance to MEM. Whole-genome sequencing revealed strain heterogeneity and different evolutionary paths, with three genes mutated in three or more strains: dacB in CZA-treated strains and oprD and ftsI in MEM-treated strains. Transcriptomic and proteomic analysis underlined heterogeneous strain response to antibiotic treatment with few commonly regulated genes and proteins. To identify genes potentially associated with antibiotic resistance, we built a machine learning model that could separate CZA- and MEM-resistant from sensitive strains based on gene expression and protein abundances. To test some of the identified associations, we performed CRISPR-Cas9 genome editing that demonstrated that mutations in dacB, ampD, and, to a lesser extent, in mexR directly affected CZA resistance. Overall, this study provides novel insights into the strain-specific molecular mechanisms regulating CZA resistance in Pseudomonas aeruginosa.IMPORTANCEPseudomonas aeruginosa is one of the most difficult-to-treat pathogens in the hospital, which often acquires resistance to multiple antibiotics. Ceftazidime-avibactam (CZA) is an essential antibiotic used to treat multidrug-resistant infections, but its resistance mechanisms are not well understood. Here we investigated the evolution of resistance to CZA and meropenem (MEM) in eight clinical bacterial isolates from patients' blood, urine, and sputum. While the rate of resistance evolution to MEM was higher than to CZA, MEM-resistant strains rarely acquired cross-resistance toward CZA. To identify changes at the genome, transcriptome, and proteome levels during antibiotic exposure, we performed multi-omics profiling of the evolved strains and confirmed the effect of several genes on antibiotic resistance with genetic engineering. Altogether, our study provides insights into the molecular response of P. aeruginosa to CZA and MEM and informs therapeutic interventions, suggesting that CZA could still be effective for patients infected with MEM-resistant pathogens.
Keywords: CRISPR/Cas9; Pseudomonas aeruginosa; ceftazidime-avibactam; dacB; genomics; machine learning; meropenem; proteomics; resistance mechanisms; transcriptomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
