The mechanism of lineage-specific tRNA recognition by bacterial tryptophanyl-tRNA synthetase and its implications for inhibitor discovery
- PMID: 40464690
- PMCID: PMC12135181
- DOI: 10.1093/nar/gkaf466
The mechanism of lineage-specific tRNA recognition by bacterial tryptophanyl-tRNA synthetase and its implications for inhibitor discovery
Abstract
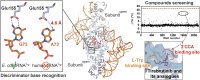
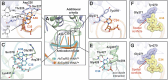
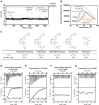
Tryptophanyl-tRNA synthetase (TrpRS) catalyzes the attachment of tryptophan (l-Trp) to tRNATrp, thereby providing the ribosome with a crucial substrate for the decoding of the UGG codon during protein translation. Both bacterial and eukaryotic TrpRSs are unable to efficiently cross-aminoacylate their respective tRNATrp substrates, indicating the evolution of lineage-specific mechanisms for tRNATrp recognition. Herein, we present the first co-crystal structure of bacterial TrpRS from Escherichia coli (EcTrpRS) in complex with its tRNATrp. EcTrpRS demonstrates bacterial-specific interactions with both the anticodon triplet and the acceptor arm of tRNATrp. Particularly, the bacterial-specific residue Glu155 forms hydrogen bonds with the discriminator base G73, thereby stabilizing it in a conformation distinct from that of A73 in the eukaryotic tRNATrp bound to human TrpRS. Through compound screening, we identified tirabrutinib and its analogues as selective inhibitors of bacterial TrpRS. These compounds occupy the l-Trp and tRNATrp CCA end binding sites of bacterial TrpRS, both of which exhibit less conservation compared to the ATP binding site between bacterial and eukaryotic TrpRSs. These findings enhance our understanding of the lineage-specific recognition of tRNATrp by bacterial TrpRS and highlight the CCA end binding site as a promising target for the future development of selective bacterial TrpRS inhibitors as potential antimicrobials.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nucleic Acids Research.
Conflict of interest statement
None declared.
Figures



Similar articles
-
Recognition by tryptophanyl-tRNA synthetases of discriminator base on tRNATrp from three biological domains.J Biol Chem. 2002 Apr 19;277(16):14343-9. doi: 10.1074/jbc.M111745200. Epub 2002 Feb 7. J Biol Chem. 2002. PMID: 11834741
-
Structure of human tryptophanyl-tRNA synthetase in complex with tRNATrp reveals the molecular basis of tRNA recognition and specificity.Nucleic Acids Res. 2006 Jun 23;34(11):3246-58. doi: 10.1093/nar/gkl441. Print 2006. Nucleic Acids Res. 2006. PMID: 16798914 Free PMC article.
-
An asymmetric structure of bacterial TrpRS supports the half-of-the-sites catalytic mechanism and facilitates antimicrobial screening.Nucleic Acids Res. 2023 May 22;51(9):4637-4649. doi: 10.1093/nar/gkad278. Nucleic Acids Res. 2023. PMID: 37070195 Free PMC article.
-
Substrate selection by aminoacyl-tRNA synthetases.Nucleic Acids Symp Ser. 1995;(33):40-2. Nucleic Acids Symp Ser. 1995. PMID: 8643392 Review.
-
Mammalian tryptophanyl-tRNA synthetases.Biochimie. 1993;75(12):1027-39. doi: 10.1016/0300-9084(93)90002-a. Biochimie. 1993. PMID: 7515282 Review.
References
MeSH terms
Substances
Grants and funding
- 22177140/National Natural Science Foundation of China
- 22477146/National Natural Science Foundation of China
- 22207133/National Natural Science Foundation of China
- 2023A1515012453/Guangdong Basic and Applied Basic Research Foundation
- 2023A1515012936/Guangdong Basic and Applied Basic Research Foundation
LinkOut - more resources
Full Text Sources

