Histone lactylation-driven feedback loop modulates cholesterol-linked immunosuppression in pancreatic cancer
- PMID: 40467104
- PMCID: PMC12573332
- DOI: 10.1136/gutjnl-2024-334361
Histone lactylation-driven feedback loop modulates cholesterol-linked immunosuppression in pancreatic cancer
Abstract
Background: Pancreatic cancer exhibits limited clinical responses to immunotherapy, highlighting the need for new strategies to counteract its immunosuppressive microenvironment. Although metabolic reprogramming and epigenetic changes contribute to malignancy, the impact of lactate-driven histone lactylation on the tumour microenvironment (TME) has not been fully explored.
Objective: This study aims to investigate the role of histone lactylation in pancreatic cancer, focusing on its effects on cholesterol metabolism and antitumour immunity.
Designs: Global lactylome profiling was conducted to identify novel epigenetic mechanisms driven by lactate-induced histone lactylation. Mechanisms were investigated via RNA sequencing, CUT&Tag, immunoprecipitation-mass spectrometry and GST-pull down. Mass cytometry by time-of-flight, in vitro co-culture system, orthotopic pancreatic cancer models and flow cytometry were used to explore Acetyl-CoA acetyltransferase (ACAT2) functions. A proteolysis-targeting chimaera (PROTAC) was developed to degrade ACAT2.
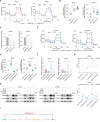
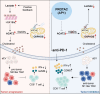
Results: Global lactylome profiling revealed that lactate-driven histone lactylation, particularly H3K18la, promotes the transcriptional activation of ACAT2. ACAT2 acetylates mitochondrial carrier homolog 2 (MTCH2), stabilising it and disrupting oxidative phosphorylation, which increases lactate production and fuels a positive feedback loop in pancreatic cancer. This loop facilitates the delivery of cholesterol via small extracellular vesicles (sEVs), polarising tumour-associated macrophages toward an immunosuppressive M2 phenotype. Additionally, the PROTAC targeting ACAT2 enhanced the efficacy of immune checkpoint blockade therapy in vivo.
Conclusions: Our findings highlight the critical role of the H3K18la/ACAT2/sEV-cholesterol axis in TME reprogramming. Targeting this pathway may improve anti-PD-1 therapy response in pancreatic cancer, providing a novel therapeutic strategy by linking histone lactylation, cholesterol metabolic reprogramming and immune modulation.
Keywords: GENE TARGETING; IMMUNE RESPONSE; LIPID METABOLISM; MACROPHAGES; PANCREATIC CANCER.
© Author(s) (or their employer(s)) 2025. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ Group.
Conflict of interest statement
Competing interests: None declared.
Figures






References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
