Matrix stiffness-induced IKBKE and MAPK8 signaling drives a phenotypic switch from DCIS to invasive breast cancer
- PMID: 40468448
- PMCID: PMC12139146
- DOI: 10.1186/s12964-025-02276-y
Matrix stiffness-induced IKBKE and MAPK8 signaling drives a phenotypic switch from DCIS to invasive breast cancer
Abstract
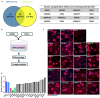
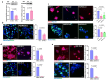
Ductal carcinoma in situ (DCIS) is not life threatening unless it transitions into invasive breast cancer (IBC). However, although breast cancer cell exposure to matrix stiffening in vitro phenotypically mimics the DCIS to IBC switch, the molecular changes driving this switch remains unclear. Here, breast cancer cell kinome activity profiling suggested matrix stiffness-upregulation of 53 kinases, among which 16 kinases were also regulated by integrin β1. Functional validation identified matrix stiffness-activation of inhibitor of nuclear factor kappa-B kinase subunit epsilon (IKBKE) and mitogen-activated protein kinase 8 (MAPK8) signaling as critical for the stiffness-driven IBC phenotype, including for cell proliferation. The IKBKE-inhibitor Amlexanox, clinically utilized for aphthous ulcers, as well as the MAPK8 inhibitor JNK-IN-8, reinstalled the DCIS-like phenotype of breast cancer cells on high matrix stiffness. This suggests that IKBKE and/or MAPK8 inhibitors could enhance the arsenal of treatments to prevent or treat breast cancer.
Keywords: Cellular signaling; Integrin; Kinase; Kinome; Mechanotransduction.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures



References
-
- Sung H, et al. Global Cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71(3):209–49. - PubMed
-
- Shapiro S. Evidence on screening for breast cancer from a randomized trial. Cancer. 1977;39(6 Suppl):2772–82. - PubMed
-
- Tabár L, et al. Reduction in mortality from breast cancer after mass screening with mammography. Randomised trial from the breast Cancer screening working group of the Swedish National board of health and welfare. Lancet. 1985;1(8433):829–32. - PubMed
-
- Ernster VL, et al. Incidence of and treatment for ductal carcinoma in situ of the breast. JAMA. 1996;275(12):913–8. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

