Genetic and functional characterization of inherited complex chromosomal rearrangements in a family with multisystem anomalies
- PMID: 40469905
- PMCID: PMC12136847
- DOI: 10.1016/j.gimo.2025.103423
Genetic and functional characterization of inherited complex chromosomal rearrangements in a family with multisystem anomalies
Abstract
Purpose: Complex chromosomal rearrangements (CCRs) are rare structural variants involving 3 or more chromosomal breakpoints. Most de novo-reported CCRs pose challenges for diagnosis and management. They often require karyotyping, fluorescence in situ hybridization, and chromosomal microarray analysis (CMA) for clinical diagnosis because of the limitations of each method. Here, we report an inherited, exceptionally complex CCR involving 4 chromosomes and 13 breakpoints in a family with multisystem anomalies.
Methods: We evaluated the CCRs using karyotyping, fluorescence in situ hybridization, CMA, and 2 emerging genomic technologies: high-throughput chromosome conformation capture sequencing aka genomic proximity mapping and optical genome mapping. We also performed functional studies using transcriptome and methylome analyses.
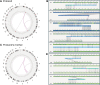
Results: The proband, who had intellectual disability and immune deficiency, shared CCRs with her unaffected mother involving chromosomes 1, 7, and 11 by karyotyping. However, CMA revealed a duplication and 3 deletions in the proband, in contrast to her mother's balanced genome. High-throughput chromosome conformation capture sequencing aka genomic proximity mapping and optical genome mapping detected the CCRs and copy-number alterations but also uncovered additional breakpoints at high resolution, including an insertion in 4p and 2 cryptic rearrangements at 7p. Transcriptome and methylome analyses identified likely biological pathways associated with the proband's phenotypes.
Conclusion: Combining cytogenetic and genomic methods provided comprehensive characterization and defined the breakpoints at high resolution in both proband and mother. This underscores the value of novel cytogenetic and genomic techniques in deciphering complex genome rearrangements and the significance of integrative genomic analysis and functional characterization in understanding clinical phenotypes.
Keywords: Complex chromosomal rearrangements; Genomic proximity mapping (GPM); Hi-C; Optical genome mapping (OGM).
© 2025 The Authors.
Conflict of interest statement
Stephen M. Eacker is an employee of Phase Genomics, Inc, the developer of the GPM technology. All other authors declare no conflicts of interest.
Figures





References
-
- Patsalis P.C. Complex chromosomal rearrangements. Genet Couns. 2007;18(1):57–69. - PubMed
LinkOut - more resources
Full Text Sources

