USP25 maintains KRAS expression and inhibiting the deubiquitinase suppresses KRAS signaling in human cancer
- PMID: 40473213
- PMCID: PMC12269508
- DOI: 10.1016/j.jbc.2025.110337
USP25 maintains KRAS expression and inhibiting the deubiquitinase suppresses KRAS signaling in human cancer
Abstract
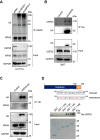
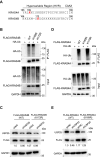
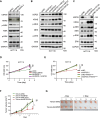
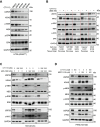
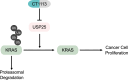
KRAS is a prominent oncogene mutated in a large number of human malignancies, particularly in pancreatic, colorectal, and lung tumors. We demonstrate here that KRAS, including its various activating mutants, is subjected to ubiquitin-mediated proteasomal degradation in cancer cells. Through an siRNA-based screening of deubiquitinases, we identified USP25 as a deubiquitinase for KRAS. Depleting USP25 expression increases ubiquitination and proteasomal degradation of KRAS, leading to the suppression of its oncogenic activity. We further show that USP25 inhibitors we have discovered are capable of destabilizing KRAS in cancer cells and are efficacious in blocking tumor xenograft growth in mice. These findings provide evidence supporting the notion that targeting the deubiquitinase USP25 can effectively, albeit indirectly, suppress KRAS and potentially aid in the treatment of tumors driven by KRAS-activating mutations.
Keywords: KRAS; USP25; proteasomal degradation; ubiquitination.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures




Similar articles
-
KRAS4B oncogenic mutants promote non-small cell lung cancer progression via the interaction of deubiquitinase USP25 with RNF31.Dev Cell. 2025 Jun 23;60(12):1768-1783.e5. doi: 10.1016/j.devcel.2025.01.015. Epub 2025 Feb 13. Dev Cell. 2025. PMID: 39952242
-
Clinicopathologic characteristics and gene expression analyses of non-KRAS 12/13, RAS-mutated metastatic colorectal cancer.Ann Oncol. 2014 Oct;25(10):2008-2014. doi: 10.1093/annonc/mdu252. Epub 2014 Jul 9. Ann Oncol. 2014. PMID: 25009008 Free PMC article.
-
USP25 directly interacts with and deubiquitinates PPARα to increase PPARα stability in hepatocytes and attenuate high-fat diet-induced MASLD in mice.Cell Death Differ. 2025 Jun;32(6):1112-1127. doi: 10.1038/s41418-025-01444-4. Epub 2025 Jan 18. Cell Death Differ. 2025. PMID: 39827322
-
KRAS mutation testing of tumours in adults with metastatic colorectal cancer: a systematic review and cost-effectiveness analysis.Health Technol Assess. 2014 Oct;18(62):1-132. doi: 10.3310/hta18620. Health Technol Assess. 2014. PMID: 25314637 Free PMC article.
-
Promising biomarkers for predicting the outcomes of patients with KRAS wild-type metastatic colorectal cancer treated with anti-epidermal growth factor receptor monoclonal antibodies: a systematic review with meta-analysis.Int J Cancer. 2013 Oct 15;133(8):1914-25. doi: 10.1002/ijc.28153. Epub 2013 Jul 13. Int J Cancer. 2013. PMID: 23494461
References
-
- Clague M.J., Urbe S. Ubiquitin: same molecule, different degradation pathways. Cell. 2010;143:682–685. - PubMed
-
- Lecker S.H., Goldberg A.L., Mitch W.E. Protein degradation by the ubiquitin-proteasome pathway in normal and disease states. J. Am. Soc. Nephrol. 2006;17:1807–1819. - PubMed
-
- Clague M.J., Barsukov I., Coulson J.M., Liu H., Rigden D.J., Urbe S. Deubiquitylases from genes to organism. Physiol. Rev. 2013;93:1289–1315. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

