Ca2+-dependent cytoplasmic and nuclear phosphorylation of STOP1 by CPK21 and CPK23 confers ALMT1-dependent aluminum resistance
- PMID: 40473618
- PMCID: PMC12141471
- DOI: 10.1038/s41467-025-60426-9
Ca2+-dependent cytoplasmic and nuclear phosphorylation of STOP1 by CPK21 and CPK23 confers ALMT1-dependent aluminum resistance
Abstract
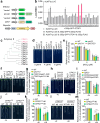
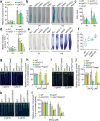
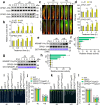
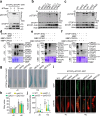
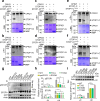
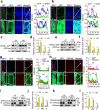
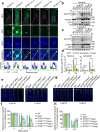
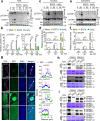
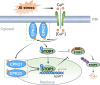
Calcium (Ca) signalling is critical for plant responses to aluminum (Al) stress, with STOP1-mediated ALMT1 expression playing a crucial role in Arabidopsis Al resistance. However, the specific intracellular Ca2+ sensors responsible for transducing Al signals in this process remain unclear. In this study, we identified CPK21 and CPK23, members of the CPK family, as key regulators promoting STOP1-mediated ALMT1 expression under Al stress, significantly influencing malate exudation from roots to limit Al accumulation in root tips. Al stress triggers rapid Ca2+-dependent accumulation of CPK21 and CPK23 in the plasma membrane, cytoplasm and nucleus of root apical cells. The Al-activated CPK21 and CPK23 subsequently phosphorylate STOP1 in both the cytoplasm and nucleus of root apical cells, stabilizing STOP1 by preventing its interaction with RAE1, ultimately enhancing Al resistance. This entire process is Ca2+-dependent. The study unveils a previously undisclosed regulatory network in which CPKs integrate Al-evoked Ca2+ signals and transcriptional reprogramming through the Ca2+-CPK21/23-STOP1 cascade to effectively respond to and adapt to Al stress in plants.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
The RAE1-STOP1-GL2-RHD6 module regulates the ALMT1-dependent aluminum resistance in Arabidopsis.Nat Commun. 2024 Jul 26;15(1):6294. doi: 10.1038/s41467-024-50784-1. Nat Commun. 2024. PMID: 39060273 Free PMC article.
-
CPK28-mediated Ca2+ signaling regulates STOP1 localization and accumulation to facilitate plant aluminum resistance.Nat Commun. 2025 Jun 5;16(1):5224. doi: 10.1038/s41467-025-60427-8. Nat Commun. 2025. PMID: 40473612 Free PMC article.
-
Under phosphate starvation conditions, Fe and Al trigger accumulation of the transcription factor STOP1 in the nucleus of Arabidopsis root cells.Plant J. 2019 Sep;99(5):937-949. doi: 10.1111/tpj.14374. Epub 2019 Jun 4. Plant J. 2019. PMID: 31034704 Free PMC article.
-
Aluminum resistance in plants: A critical review focusing on STOP1.Plant Commun. 2025 Feb 10;6(2):101200. doi: 10.1016/j.xplc.2024.101200. Epub 2024 Dec 2. Plant Commun. 2025. PMID: 39628052 Free PMC article. Review.
-
Synergistic and antagonistic pleiotropy of STOP1 in stress tolerance.Trends Plant Sci. 2021 Oct;26(10):1014-1022. doi: 10.1016/j.tplants.2021.06.011. Epub 2021 Jul 9. Trends Plant Sci. 2021. PMID: 34253485 Review.
References
-
- von Uexkull, H. R. & Mutert, E. Global extent, development and economic-impact of acid soils. Plant Soil171, 1–15 (1995).
-
- Horst, W. J., Asher, C. J., Cakmak, J., Szulkiewicz, P. & Wissemeier, A. H. Short-term responses of soybean roots to aluminium. J. Plant Physiol.140, 174–178 (1992).
-
- Kochian, L. V., Pineros, M. A., Liu, J. & Magalhaes, J. V. Plant adaptation to acid soils: the molecular basis for crop aluminum resistance. Annu. Rev. Plant Biol.66, 571–598 (2015). - PubMed
-
- Ma, J. F., Ryan, P. R. & Delhaize, E. Aluminium tolerance in plants and the complexing role of organic acids. Trends Plant Sci.6, 273–278 (2001). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

