Model systems and unique biological features of high and low-grade colorectal cancer (CRC) revealed by xenografting 84 human CRC cell lines
- PMID: 40473896
- PMCID: PMC12141688
- DOI: 10.1038/s42003-025-08251-0
Model systems and unique biological features of high and low-grade colorectal cancer (CRC) revealed by xenografting 84 human CRC cell lines
Abstract
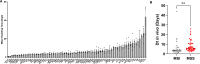
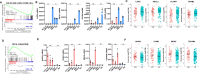
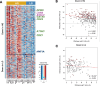
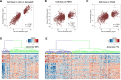
Colorectal cancers (CRCs) present across a range of differentiation grades, which impact patient outcome and management; however, the molecular features and drivers of differentiation status are not fully understood. To address this, 84 commonly used human CRC cell lines were grown as xenografts in mice, revealing models of low-grade (LG) and high-grade (HG) CRC. Transcriptional profiling revealed coordinate downregulation of multiple transcription factors involved in intestinal development and differentiation, markers of colonic lineage-specific differentiation, and effectors of normal functions of the colonic epithelium in HG tumours. Mechanistically, multiple genes suppressed in HG tumours harboured promoter methylation, indicative of stable epigenetic silencing. Furthermore, markers of LGR5+ colon stem cells were suppressed in HG tumours, while markers of cell proliferation, fetal-like intestinal stem cells, and non-canonical cell types including mesenchymal cells were increased. These changes manifested in HG cell line displaying increased proliferation, migration and metastatic capacity. Importantly, CRC cell line-derived transcriptional profiles of differentiation grade were reflected in LG and HG patient-derived tumour organoids and primary CRCs, revealing cell lines accurately model differentiation grade. The models and tumour differentiation-related properties identified herein may inform new approaches for tailored CRC treatments based on tumour grade.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

