Enterobacterales use capsules, transporters, mobile genetic elements, and other evolutionary adaptations to survive antibiotics exposure in the absence of resistance genes
- PMID: 40474678
- PMCID: PMC12147488
- DOI: 10.1080/21505594.2025.2514092
Enterobacterales use capsules, transporters, mobile genetic elements, and other evolutionary adaptations to survive antibiotics exposure in the absence of resistance genes
Abstract
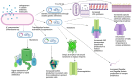
Methods: Whole-genome sequencing, transcriptomic profiling, and epigenomic analyses were performed. Phenotypic assays were used to evaluate the effects of various inhibitors on antibiotic susceptibility, while bioinformatic pipelines were used to characterize resistance determinants, virulence factors, and mobile genetic elements (MGEs).
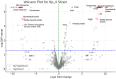
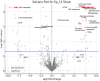
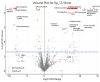
Results: Phylogenetic analysis revealed widespread carriage of diverse resistance genes, particularly on plasmids of K. pneumoniae, while Enterobacter species possessed fewer known ARGs. Despite lacking known carbapenemase and mcr genes, several isolates demonstrated colistin or carbapenem resistance mediated by upregulation of efflux pumps, overproduction of capsular polysaccharides, mutations in outer membrane proteins, and potential lipopolysaccharide-modifying enzymes. Transcriptomic analysis revealed significant differential gene expression upon antibiotic exposure. Notably, genes encoding ABC transporter proteins were significantly downregulated (p-value <0.0001, fold change > 10), while genes encoding transposases were significantly upregulated (p-value <0.0001, fold change > 11). These changes underscore the critical role of transporters and MGEs in antibiotic resistance adaptation.
Conclusions: In the absence of canonical carbapenemase and mcr genes, K. pneumoniae and Enterobacter species can deploy a spectrum of adaptive mechanisms, including efflux pumps, mobile elements, and altered outer membrane/capsule structures, to overcome colistin and carbapenem treatments. These findings support the need for ongoing surveillance of novel or underrecognized resistance mechanisms to preserve the efficacy of last-line antibiotics.
Keywords: Multi-drug resistance; RNA-sequencing; epigenomics; genomics; transcriptomic profiling.
Conflict of interest statement
No potential conflict of interest was reported by the author(s).
Figures





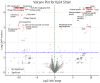


References
-
- Mmatli M, Leshaba TMS, Skosana LB, et al. Molecular screening of clinical multidrug-resistant Gram-negative bacteria shows endemicity of carbapenemases, coexistence of multiple carbapenemases, and rarity of mcr in South Africa. Microb Drug Resist. 2022;28(11):1028–1036. doi: 10.1089/mdr.2022.0112 - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
