Upstream open reading frames buffer translational variability during Drosophila evolution and development
- PMID: 40478227
- PMCID: PMC12143884
- DOI: 10.7554/eLife.104074
Upstream open reading frames buffer translational variability during Drosophila evolution and development
Abstract
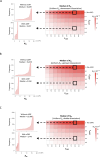
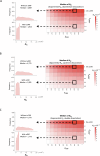
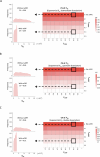
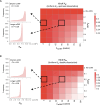
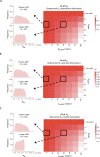
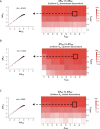
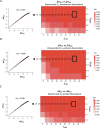
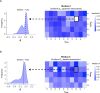
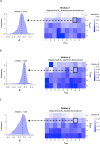
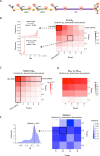
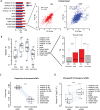
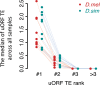
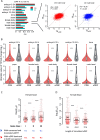
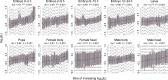
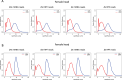
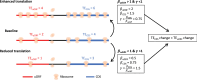
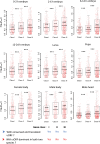
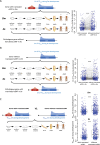
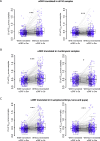
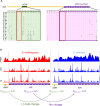
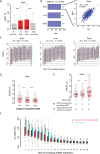
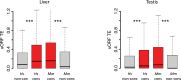
Protein abundance tends to be more evolutionarily conserved than mRNA levels both within and between species, yet the mechanisms underlying this phenomenon remain largely unknown. Upstream open reading frames (uORFs) are widespread cis-regulatory elements in eukaryotic genomes that regulate translation, but it remains unclear whether and how uORFs contribute to stabilizing protein levels. In this study, we performed ribosome translation simulations on mRNA to quantitatively assess the extent to which uORF translation influences the translational variability of downstream coding sequences (CDSs) across varying contexts. Our simulations revealed that uORF translation dampens CDS translational variability, with buffering capacity increasing in proportion to uORF translation efficiency, length, and number. We then compared the translatomes at different developmental stages of two Drosophila species, demonstrating that uORFs buffer mRNA translation fluctuations during both evolution and development. Experimentally, deleting a uORF in the bicoid (bcd) gene-a prominent example of translational buffering-resulted in extensive changes in gene expression and phenotypes in Drosophila melanogaster. Additionally, we observed uORF-mediated buffering between primates and within human populations. Together, our results reveal a novel regulatory mechanism by which uORFs stabilize gene translation during development and across evolutionary time.
Keywords: TASEP modeling; bicoid; drosophila melanogaster; drosophila simulans; evolutionary biology; genetics; genomics; humans; translation evolution; translational regulation; uORF.
© 2025, Sun, Duan et al.
Conflict of interest statement
YS, YD, PG, CL, KJ, SD, WT, HZ, JL No competing interests declared
Figures
















Update of
- doi: 10.1101/2024.11.13.623404
- doi: 10.7554/eLife.104074.1
- doi: 10.7554/eLife.104074.2
Similar articles
-
Ribo-uORF: a comprehensive data resource of upstream open reading frames (uORFs) based on ribosome profiling.Nucleic Acids Res. 2023 Jan 6;51(D1):D248-D261. doi: 10.1093/nar/gkac1094. Nucleic Acids Res. 2023. PMID: 36440758 Free PMC article.
-
Translational buffering by ribosome stalling in upstream open reading frames.PLoS Genet. 2022 Oct 31;18(10):e1010460. doi: 10.1371/journal.pgen.1010460. eCollection 2022 Oct. PLoS Genet. 2022. PMID: 36315596 Free PMC article.
-
Genome-wide maps of ribosomal occupancy provide insights into adaptive evolution and regulatory roles of uORFs during Drosophila development.PLoS Biol. 2018 Jul 20;16(7):e2003903. doi: 10.1371/journal.pbio.2003903. eCollection 2018 Jul. PLoS Biol. 2018. PMID: 30028832 Free PMC article.
-
Regulation of plant translation by upstream open reading frames.Plant Sci. 2014 Jan;214:1-12. doi: 10.1016/j.plantsci.2013.09.006. Epub 2013 Sep 18. Plant Sci. 2014. PMID: 24268158 Review.
-
Function and Evolution of Upstream ORFs in Eukaryotes.Trends Biochem Sci. 2019 Sep;44(9):782-794. doi: 10.1016/j.tibs.2019.03.002. Epub 2019 Apr 16. Trends Biochem Sci. 2019. PMID: 31003826 Review.
References
-
- Amourda C, Chong J, Saunders TE. MicroRNAs buffer genetic variation at specific temperatures during embryonic development. bioRxiv. 2018 doi: 10.1101/444810. - DOI
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

