A Knowledge-Enhanced Platform (MetaSepsisKnowHub) for Retrieval Augmented Generation-Based Sepsis Heterogeneity and Personalized Management: Development Study
- PMID: 40478618
- PMCID: PMC12181755
- DOI: 10.2196/67201
A Knowledge-Enhanced Platform (MetaSepsisKnowHub) for Retrieval Augmented Generation-Based Sepsis Heterogeneity and Personalized Management: Development Study
Abstract
Background: Sepsis is a severe syndrome of organ dysfunction caused by infection; it has high heterogeneity and high in-hospital mortality, representing a grim clinical challenge for precision medicine in critical care.
Objective: We aimed to extract reported sepsis biomarkers to provide users with comprehensive biomedical information and integrate retrieval augmented generation (RAG) and prompt engineering to enhance the accuracy, stability, and interpretability of clinical decisions recommended by large language models (LLMs).
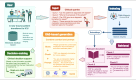
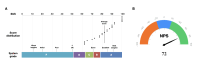
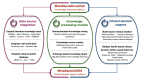
Methods: To address the challenge, we established and updated the first knowledge-enhanced platform, MetaSepsisKnowHub, comprising 427 sepsis biomarkers and 423 studies, aiming to systematically collect and annotate sepsis biomarkers to guide personalized clinical decision-making in the diagnosis and treatment of human sepsis. We curated a tailored LLM framework incorporating RAG and prompt engineering and incorporated 2 performance evaluation scales: the System Usability Scale and the Net Promoter Score.
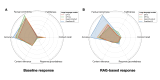
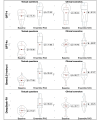
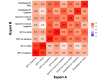
Results: The overall quantitative ratings of expert-reviewed clinical recommendations based on RAG surpassed baseline responses generated by 4 LLMs and showed a statistically significant improvement in textual questions (GPT-4: mean 75.79, SD 7.11 vs mean 81.59, SD 9.87; P=.02; GPT-4o: mean 70.36, SD 7.63 vs mean 77.98, SD 13.26; P=.02; Qwen2.5-instruct: mean 77.08 SD 3.75 vs mean 85.46, SD 7.27; P<.001; and DeepSeek-R1: mean 77.67, SD 3.66 vs mean 86.42, SD 8.56; P<.001), but no significant statistical differences could be measured in clinical scenarios. The RAG assessment score comparing RAG-based responses and expert-provided benchmark answers illustrated prominent factual correctness, accuracy, and knowledge recall compared to the baseline responses. After use, the average the System Usability Scale score was 82.20 (SD 14.17) and the Net Promoter Score was 72, demonstrating high user satisfaction and loyalty.
Conclusions: We highlight the pioneering MetaSepsisKnowHub platform, and we show that combining MetaSepsisKnowHub with RAG can minimize limitations on precision and maximize the breadth of LLMs to shorten the bench-to-bedside distance, serving as a knowledge-enhanced paradigm for future application of artificial intelligence in critical care medicine.
Keywords: human sepsis; knowledge-enhanced; personalized application; precision medicine; retrieval augmented generation.
©Chi Zhang, Hao Yang, Xingyun Liu, Rongrong Wu, Hui Zong, Erman Wu, Yi Zhou, Jiakun Li, Bairong Shen. Originally published in the Journal of Medical Internet Research (https://www.jmir.org), 06.06.2025.
Conflict of interest statement
Conflicts of Interest: None declared.
Figures



References
-
- Singer M, Deutschman CS, Seymour CW, Shankar-Hari M, Annane D, Bauer M, Bellomo R, Bernard GR, Chiche JD, Coopersmith CM, Hotchkiss RS, Levy MM, Marshall JC, Martin GS, Opal SM, Rubenfeld GD, van der Poll T, Vincent JL, Angus DC. The third international consensus definitions for sepsis and septic shock (sepsis-3) JAMA. 2016 Feb 23;315(8):801–10. doi: 10.1001/jama.2016.0287. https://europepmc.org/abstract/MED/26903338 2492881 - DOI - PMC - PubMed
-
- Evans L, Rhodes A, Alhazzani W, Antonelli M, Coopersmith CM, French C, Machado FR, Mcintyre L, Ostermann M, Prescott HC, Schorr C, Simpson S, Wiersinga WJ, Alshamsi F, Angus DC, Arabi Y, Azevedo L, Beale R, Beilman G, Belley-Cote E, Burry L, Cecconi M, Centofanti J, Coz Yataco A, De Waele J, Dellinger RP, Doi K, Du B, Estenssoro E, Ferrer R, Gomersall C, Hodgson C, Møller MH, Iwashyna T, Jacob S, Kleinpell R, Klompas M, Koh Y, Kumar A, Kwizera A, Lobo S, Masur H, McGloughlin S, Mehta S, Mehta Y, Mer M, Nunnally M, Oczkowski S, Osborn T, Papathanassoglou E, Perner A, Puskarich M, Roberts J, Schweickert W, Seckel M, Sevransky J, Sprung CL, Welte T, Zimmerman J, Levy M. Surviving sepsis campaign: international guidelines for management of sepsis and septic shock 2021. Intensive Care Med. 2021 Nov;47(11):1181–247. doi: 10.1007/s00134-021-06506-y. https://europepmc.org/abstract/MED/34599691 10.1007/s00134-021-06506-y - DOI - PMC - PubMed
-
- Freund Yonathan, Lemachatti Najla, Krastinova Evguenia, Van Laer Marie, Claessens Yann-Erick, Avondo Aurélie, Occelli Céline, Feral-Pierssens Anne-Laure, Truchot Jennifer, Ortega Mar, Carneiro Bruno, Pernet Julie, Claret Pierre-Géraud, Dami Fabrice, Bloom Ben, Riou Bruno, Beaune Sébastien, French Society of Emergency Medicine Collaborators Group Prognostic accuracy of sepsis-3 criteria for in-hospital mortality among patients with suspected infection presenting to the emergency department. JAMA. 2017 Jan 17;317(3):301–308. doi: 10.1001/jama.2016.20329. https://core.ac.uk/reader/84058949?utm_source=linkout 2598268 - DOI - PubMed
-
- Rudd KE, Johnson SC, Agesa KM, Shackelford KA, Tsoi D, Kievlan DR, Colombara DV, Ikuta KS, Kissoon N, Finfer S, Fleischmann-Struzek C, Machado FR, Reinhart KK, Rowan K, Seymour CW, Watson RS, West TE, Marinho F, Hay SI, Lozano R, Lopez AD, Angus DC, Murray CJ, Naghavi M. Global, regional, and national sepsis incidence and mortality, 1990-2017: analysis for the Global Burden of Disease Study. Lancet. 2020 Jan 18;395(10219):200–211. doi: 10.1016/S0140-6736(19)32989-7. https://linkinghub.elsevier.com/retrieve/pii/S0140-6736(19)32989-7 S0140-6736(19)32989-7 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

