Chromosome-level reference genome for the medically important Arabian horned viper (Cerastes gasperettii)
- PMID: 40478730
- PMCID: PMC12143202
- DOI: 10.1093/gigascience/giaf030
Chromosome-level reference genome for the medically important Arabian horned viper (Cerastes gasperettii)
Abstract
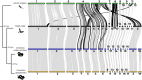
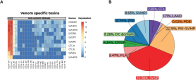
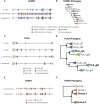
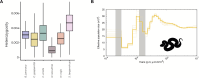
Venoms have traditionally been studied from a proteomic and/or transcriptomic perspective, often overlooking the true genetic complexity underlying venom production. The recent surge in genome-based venom research (sometimes called "venomics") has proven to be instrumental in deepening our understanding of venom evolution at the molecular level, particularly through the identification and mapping of toxin-coding loci across the broader chromosomal architecture. Although venomous snakes are a model system in venom research, the number of high-quality reference genomes in the group remains limited. In this study, we present a chromosome-resolution reference genome for the Arabian horned viper Cerastes gasperettii (NCBI: txid110202), a venomous snake native to the Arabian Peninsula. Our highly contiguous genome (genome size: 1.63 Gbp; contig N50: 45.6 Mbp; BUSCO: 92.8%) allowed us to explore macrochromosomal rearrangements within the Viperidae family, as well as across squamates. We identified the main highly expressed toxin genes within the venom glands comprising the venom's core, in line with our proteomic results. We also compared microsyntenic changes in the main toxin gene clusters with those of other venomous snake species, highlighting the pivotal role of gene duplication and loss in the emergence and diversification of snake venom metalloproteinases and snake venom serine proteases for C. gasperettii. Using Illumina short-read sequencing data, we reconstructed the demographic history and genome-wide heterozigosity of the species, revealing how historical aridity likely drove population expansions. Finally, this study highlights the importance of using long-read sequencing as well as chromosome-level reference genomes to disentangle the origin and diversification of toxin gene families in venomous snake species.
Keywords: gene synteny; genomics; toxin evolution; transcriptomics; venom.
© The Author(s) 2025. Published by Oxford University Press GigaScience.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Successful management of severe coagulopathy in a dog following Cerastes gasperettii (Arabian horned Viper) envenomation: First clinical case description in Israel.Toxicon. 2025 Apr;258:108321. doi: 10.1016/j.toxicon.2025.108321. Epub 2025 Mar 10. Toxicon. 2025. PMID: 40074044
-
Hidden in the sand: Phylogenomics unravel an unexpected evolutionary history for the desert-adapted vipers of the genus Cerastes.Mol Phylogenet Evol. 2024 Feb;191:107979. doi: 10.1016/j.ympev.2023.107979. Epub 2023 Nov 29. Mol Phylogenet Evol. 2024. PMID: 38040070
-
Comprehensive Study of the Proteome and Transcriptome of the Venom of the Most Venomous European Viper: Discovery of a New Subclass of Ancestral Snake Venom Metalloproteinase Precursor-Derived Proteins.J Proteome Res. 2019 May 3;18(5):2287-2309. doi: 10.1021/acs.jproteome.9b00120. Epub 2019 Apr 24. J Proteome Res. 2019. PMID: 31017792 Free PMC article.
-
Venomous snakes of Costa Rica: biological and medical implications of their venom proteomic profiles analyzed through the strategy of snake venomics.J Proteomics. 2014 Jun 13;105:323-39. doi: 10.1016/j.jprot.2014.02.020. Epub 2014 Feb 24. J Proteomics. 2014. PMID: 24576642 Review.
-
A Review of the Proteomic Profiling of African Viperidae and Elapidae Snake Venoms and Their Antivenom Neutralisation.Toxins (Basel). 2022 Oct 22;14(11):723. doi: 10.3390/toxins14110723. Toxins (Basel). 2022. PMID: 36355973 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

