Spatio-temporal organization of network activity patterns in the hippocampus
- PMID: 40478735
- PMCID: PMC7617751
- DOI: 10.1016/j.celrep.2025.115808
Spatio-temporal organization of network activity patterns in the hippocampus
Abstract
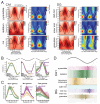
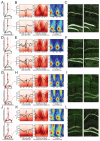
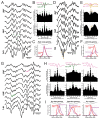
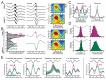
Understanding how coordinated neural networks support brain functions remains a central goal in neuroscience. The hippocampus, with its layered architecture and structured inputs to diverse cell populations, is a tractable model for dissecting operating microcircuits through the analysis of electrophysiological signatures. We investigated hippocampal network patterns in behaving mice by developing a low-dimensional embedding of local field potentials recorded along the CA1-to-dentate gyrus axis. This embedding revealed layer-specific gamma profiles reflecting spatially organized rhythms and their associated principal cell-interneuron firing motifs. Moreover, firing behaviors along the CA1 radial axis distinguished between deep and superficial principal cells, as well as between interneurons from the pyramidal, radiatum, and lacunosum-moleculare layers. These findings provide a comprehensive map of spatiotemporal activity patterns underlying hippocampal network functions.
Keywords: CP: Neuroscience; firing motifs; hippocampus; network activity; neuronal populations; oscillations.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures



References
-
- Andersen P, editor. The hippocampus book. Oxford University Press; 2007.
-
- Eichenbaum H. A cortical–hippocampal system for declarative memory. Nat Rev Neurosci. 2000;1:41–50. - PubMed
-
- Somogyi P, Klausberger T. In: Handbook of Brain Microcircuits. Shepherd GM, Grillner S, editors. Oxford University Press; 2017. Hippocampus: Intrinsic Organization; pp. 199–216. - DOI
-
- Van Strien NM, Cappaert NLM, Witter MP. The anatomy of memory: an interactive overview of the parahippocampal–hippocampal network. Nat Rev Neurosci. 2009;10:272–282. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

