A spatially organized Cd24a+/ Pax9+ stem cell core governs postnatal tooth establishment
- PMID: 40479070
- PMCID: PMC12143388
- DOI: 10.1126/sciadv.adu5653
A spatially organized Cd24a+/ Pax9+ stem cell core governs postnatal tooth establishment
Abstract
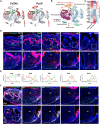
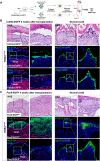
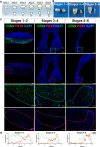
Mineralized tissues are fundamental in mechanical support and protection in vertebrates. Their formation by progenitor cells during development remains elusive. Here, we show that the postnatal establishment of the molar teeth was governed by a spatially organized core of Cd24a+/Pax9+ progenitors that persisted into adulthood. Cd24a+ cells gave rise to the dentin-pulp complex, while Pax9+ ones mainly generated periodontal tissues. During development, guided by alveolar bone-derived PDGFB (platelet-derived growth factor, B polypeptide), Cd24a+/Pax9+ cells gradually concentrated on the apical region during the crown-to-root transition, collectively migrated and formed dental root. Cell ablation and conditional Wnt knockout notably compromised tooth establishment. Single-cell RNA sequencing, CUT&Tag, and spatial mapping further revealed distinct features of Cd24a+/Pax9+ cells and their cellular organization. Last, the CD24+/PAX9+ core was also present in human teeth, suggesting it as a conserved developmental program. Together, our work underscores the role of spatially organized dental stem cells in the postnatal establishment of a model mineralized organ in mammals.
Figures






References
-
- Lowenstam H. A., Minerals formed by organisms. Science 211, 1126–1131 (1981). - PubMed
-
- Mann S., Molecular recognition in biomineralization. Nature 332, 119–124 (1988).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

