Spatiotemporal Tracking of Three Novel Transposable Element Invasions in Drosophila melanogaster over the Last 30 Years
- PMID: 40479505
- PMCID: PMC12230796
- DOI: 10.1093/molbev/msaf143
Spatiotemporal Tracking of Three Novel Transposable Element Invasions in Drosophila melanogaster over the Last 30 Years
Abstract
Transposable elements (TEs) are repetitive sequences capable of mobilizing within genomes, exerting a significant influence on evolution throughout the tree of life. Using a novel approach that does not require prior knowledge of the sequence of repeats, we identified three novel TE invasions in Drosophila melanogaster: McLE spread between 1990-2000, Souslik between 2009-2012, and Transib1 between 2013-2016. We recapitulate previous findings, revealing that a total of 11 TEs invaded D. melanogaster over the past two centuries. These 11 invasions increased the fly genome by ∼1 Mbp. Using data from over 1,400 arthropod genomes, we provide evidence that these TE invasions were triggered by horizontal transfers, with Drosophila simulans and species of the Drosophila willistoni group acting as putative donors. Through the analysis of ∼600 short-read datasets spanning diverse geographic regions, we reveal the rapidity of TE invasions: Transib1 swiftly multiplied from three isolated epicenters in 2014 to all investigated populations in just 2 years. Our findings suggest that anthropogenic activities, which facilitate the range and population expansions of D. melanogaster, could have accelerated the rate of horizontal transposon transfer as well as the spread of the TEs into the worldwide population. Given the significant impact of TEs on evolution and the potential involvement of humans in their dispersal, our research has crucial implications for both evolution and ecology.
Keywords: Drosophila; genome evolution; horizontal gene transfer; transposable elements; transposon invasions.
© The Author(s) 2025. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Conflict of interest statement
Conflicts of Interest: The author(s) declare(s) that there is no conflict of interest regarding the publication of this article.
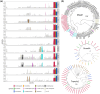
Figures



Similar articles
-
Double trouble: two retrotransposons triggered a cascade of invasions in Drosophila species within the last 50 years.Nat Commun. 2025 Jan 9;16(1):516. doi: 10.1038/s41467-024-55779-6. Nat Commun. 2025. PMID: 39788974 Free PMC article.
-
Patterns of piRNA Regulation in Drosophila Revealed through Transposable Element Clade Inference.Mol Biol Evol. 2022 Jan 7;39(1):msab336. doi: 10.1093/molbev/msab336. Mol Biol Evol. 2022. PMID: 34921315 Free PMC article.
-
Purifying selection shapes the dynamics of P-element invasion in Drosophila simulans populations.Genome Biol. 2025 Jul 24;26(1):221. doi: 10.1186/s13059-025-03688-2. Genome Biol. 2025. PMID: 40708032 Free PMC article.
-
Factors that influence parents' and informal caregivers' views and practices regarding routine childhood vaccination: a qualitative evidence synthesis.Cochrane Database Syst Rev. 2021 Oct 27;10(10):CD013265. doi: 10.1002/14651858.CD013265.pub2. Cochrane Database Syst Rev. 2021. PMID: 34706066 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2017 Dec 22;12(12):CD011535. doi: 10.1002/14651858.CD011535.pub2. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2020 Jan 9;1:CD011535. doi: 10.1002/14651858.CD011535.pub3. PMID: 29271481 Free PMC article. Updated.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

