Cryo-EM uncovers a sequential mechanism for RNA polymerase I pausing and stalling at abasic DNA lesions
- PMID: 40480971
- PMCID: PMC12144180
- DOI: 10.1038/s41467-025-60536-4
Cryo-EM uncovers a sequential mechanism for RNA polymerase I pausing and stalling at abasic DNA lesions
Abstract
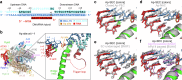
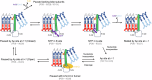
During synthesis of the ribosomal RNA precursor, RNA polymerase I (Pol I) monitors DNA integrity but its response to DNA damage remains poorly studied. Abasic sites are among the most prevalent DNA lesions in eukaryotic cells, and their detection is critical for cell survival. We report cryo-EM structures of Pol I in different stages of stalling at abasic sites, supported by in vitro transcription studies. Slow nucleotide addition opposite abasic sites occurs through base sandwiching between the RNA 3'-end and the Pol I bridge helix. Templating abasic sites can also cause Pol I cleft opening, which enables the A12 subunit to access the active center. Nucleotide addition opposite the lesion induces a translocation intermediate where DNA bases tilt to form hydrogen bonds with the new RNA base. These findings reveal unique mechanisms of Pol I stalling at abasic sites, differing from arrest by bulky lesions or abasic site handling by RNA polymerase II.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interest.
Figures





References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

