Population-scale cross-disorder atlas of the human prefrontal cortex at single-cell resolution
- PMID: 40480991
- PMCID: PMC12144096
- DOI: 10.1038/s41597-025-04687-5
Population-scale cross-disorder atlas of the human prefrontal cortex at single-cell resolution
Abstract
Neurodegenerative diseases and serious mental illnesses often exhibit overlapping characteristics, highlighting the potential for shared underlying mechanisms. To facilitate a deeper understanding of these diseases and pave the way for more effective treatments, we have generated a population-scale multi-omics dataset consisting of genotype and single-nucleus transcriptome data from the prefrontal cortex of frozen human brain specimens. Encompassing over 6.3 million nuclei from 1,494 donors, our dataset represents a diverse range of neurodegenerative and serious mental illnesses, including Alzheimer's and Parkinson's diseases, schizophrenia, bipolar disorder and diffuse Lewy body dementia, as well as neurotypical controls. Our dataset offers a unique opportunity to study disease interactions, as 21% of donors had comorbid diagnoses of two or more major brain disorders. Additionally, it includes detailed phenotypic information on neuropsychiatric symptoms, such as apathy and weight loss, which commonly accompany Alzheimer's disease and related dementias. We have performed stringent preprocessing and quality controls, ensuring the reliability and usability of the data. As a commitment to fostering collaborative research, we provide this valuable resource as an online repository, enabling widespread analyses across the scientific community.
© 2025. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
Competing interests: Panos Roussos is an Editorial Board Member for Scientific Data.
Figures

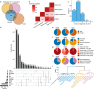
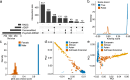


References
Publication types
MeSH terms
Grants and funding
- R01 AG067025/AG/NIA NIH HHS/United States
- R01AG065582/U.S. Department of Health & Human Services | NIH | National Institute on Aging (U.S. National Institute on Aging)
- R01AG067025/U.S. Department of Health & Human Services | NIH | National Institute on Aging (U.S. National Institute on Aging)
- R01AG082185/U.S. Department of Health & Human Services | NIH | National Institute on Aging (U.S. National Institute on Aging)
LinkOut - more resources
Full Text Sources
Medical
Research Materials

