Investigating methods to enhance interpretability and performance in cardiac MRI for myocardial scarring diagnosis using convolutional neural network classification and One Match
- PMID: 40489519
- PMCID: PMC12148160
- DOI: 10.1371/journal.pone.0313971
Investigating methods to enhance interpretability and performance in cardiac MRI for myocardial scarring diagnosis using convolutional neural network classification and One Match
Abstract
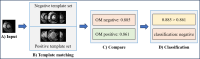
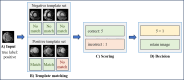
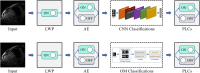
Machine learning (ML) classification of myocardial scarring in cardiac MRI is often hindered by limited explainability, particularly with convolutional neural networks (CNNs). To address this, we developed One Match (OM), an algorithm that builds on template matching to improve on both the explainability and performance of ML myocardial scaring classification. By incorporating OM, we aim to foster trust in AI models for medical diagnostics and demonstrate that improved interpretability does not have to compromise classification accuracy. Using a cardiac MRI dataset from 279 patients, this study evaluates One Match, which classifies myocardial scarring in images by matching each image to a set of labeled template images. It uses the highest correlation score from these matches for classification and is compared to a traditional sequential CNN. Enhancements such as autodidactic enhancement (AE) and patient-level classifications (PLCs) were applied to improve the predictive accuracy of both methods. Results are reported as follows: accuracy, sensitivity, specificity, precision, and F1-score. The highest classification performance was observed with the OM algorithm when enhanced by both AE and PLCs, 95.3% accuracy, 92.3% sensitivity, 96.7% specificity, 92.3% precision, and 92.3% F1-score, marking a significant improvement over the base configurations. AE alone had a positive impact on OM increasing accuracy from 89.0% to 93.2%, but decreased the accuracy of the CNN from 85.3% to 82.9%. In contrast, PLCs improved accuracy for both the CNN and OM, raising the CNN's accuracy by 4.2% and OM's by 7.4%. This study demonstrates the effectiveness of OM in classifying myocardial scars, particularly when enhanced with AE and PLCs. The interpretability of OM also enabled the examination of misclassifications, providing insights that could accelerate development and foster greater trust among clinical stakeholders.
Copyright: © 2025 Udin et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




References
-
- Goldberger JJ, Cain ME, Hohnloser SH, Kadish AH, Knight BP, Lauer MS, et al. American Heart Association/American College of Cardiology Foundation/Heart Rhythm Society scientific statement on noninvasive risk stratification techniques for identifying patients at risk for sudden cardiac death. A scientific statement from the American Heart Association Council on clinical Cardiology Committee on Electrocardiography and Arrhythmias and Council on Epidemiology and Prevention. J Am Coll Cardiol. 2008;52(14):1179–99. doi: 10.1016/j.jacc.2008.05.003 - DOI - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

