P53-Induced Autophagy Degradation of NKX3-2 Improves Ovarian Cancer Prognosis
- PMID: 40497941
- PMCID: PMC12153924
- DOI: 10.3390/cells14110765
P53-Induced Autophagy Degradation of NKX3-2 Improves Ovarian Cancer Prognosis
Abstract
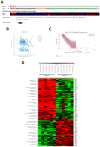
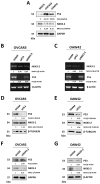
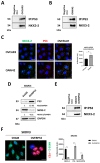
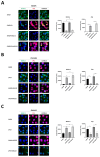
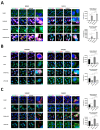
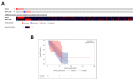
NKX3-2, a transcriptional repressor factor belonging to the NK family of homeobox-containing proteins, has been widely studied for its role in promoting chondrogenic differentiation and homeostasis. NKX3-2 is upregulated in chemoresistant ovarian tumors and metastatic gastric cancer cells; however, its prognostic role and mechanistic involvement in cancer cell biology remain to be elucidated. By interrogating the TCGA database, we found that cancer patients with high NKX3-2 expression had a shorter overall survival rate than patients with low expression. In ovarian cancer patients, NKX3-2 negatively correlates with P53. Given the prominent role of the latter oncosuppressor in controlling DNA repair and cell death, here we investigate the molecular mechanism involved in this negative correlation in several ovarian cancer cell lines expressing different levels of the two proteins. We found that the high expression of endogenous or ectopic P53 reduced NKX3-2 protein expression, while its knockdown increased it. In contrast, the genetic manipulation of NKX3-2 expression did not affect P53 expression. Mechanistically, P53-mediated downregulation of NKX3-2 does not entail transcriptional activity or proteasomal clearance but occurs via P53-NKX3-2 protein-protein interaction, which in turn results in P53-induced NKX3-2 degradation via the autophagy-lysosome pathway. Remarkably, patients bearing a tumor characterized by low NKX3-2 and high MAP1LC3B expression (indicative of active autophagy) display a better prognosis. Taken together, our data indicate that NKX3-2 represents a negative prognostic factor under P53 control in ovarian cancer. From a translational point of view, identifying this novel mechanism may represent a new molecular signature capable of predicting the clinical outcome of patients, a crucial aspect of developing personalized therapeutic approaches.
Keywords: BAPX1; TCGA; apoptosis; autophagy; cancer cell metabolism; lysosome; ovarian cancer; p53; personalized medicine; prognosis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Ouyang S., Zhu G., Ouyang L., Luo Y., Zhou R., Pan C., Bin J., Liao Y., Liao W. Bapx1 mediates transforming growth factor-β- induced epithelial-mesenchymal transition and promotes a malignancy phenotype of gastric cancer cells. Biochem. Biophys. Res. Commun. 2017;486:285–292. doi: 10.1016/j.bbrc.2017.03.029. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

