Enhancing Pulmonary Disease Prediction Using Large Language Models With Feature Summarization and Hybrid Retrieval-Augmented Generation: Multicenter Methodological Study Based on Radiology Report
- PMID: 40499132
- PMCID: PMC12176309
- DOI: 10.2196/72638
Enhancing Pulmonary Disease Prediction Using Large Language Models With Feature Summarization and Hybrid Retrieval-Augmented Generation: Multicenter Methodological Study Based on Radiology Report
Abstract
Background: The rapid advancements in natural language processing, particularly the development of large language models (LLMs), have opened new avenues for managing complex clinical text data. However, the inherent complexity and specificity of medical texts present significant challenges for the practical application of prompt engineering in diagnostic tasks.
Objective: This paper explores LLMs with new prompt engineering technology to enhance model interpretability and improve the prediction performance of pulmonary disease based on a traditional deep learning model.
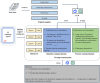
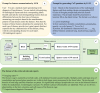
Methods: A retrospective dataset including 2965 chest CT radiology reports was constructed. The reports were from 4 cohorts, namely, healthy individuals and patients with pulmonary tuberculosis, lung cancer, and pneumonia. Then, a novel prompt engineering strategy that integrates feature summarization (F-Sum), chain of thought (CoT) reasoning, and a hybrid retrieval-augmented generation (RAG) framework was proposed. A feature summarization approach, leveraging term frequency-inverse document frequency (TF-IDF) and K-means clustering, was used to extract and distill key radiological findings related to 3 diseases. Simultaneously, the hybrid RAG framework combined dense and sparse vector representations to enhance LLMs' comprehension of disease-related text. In total, 3 state-of-the-art LLMs, GLM-4-Plus, GLM-4-air (Zhipu AI), and GPT-4o (OpenAI), were integrated with the prompt strategy to evaluate the efficiency in recognizing pneumonia, tuberculosis, and lung cancer. The traditional deep learning model, BERT (Bidirectional Encoder Representations from Transformers), was also compared to assess the superiority of LLMs. Finally, the proposed method was tested on an external validation dataset consisted of 343 chest computed tomography (CT) report from another hospital.
Results: Compared with BERT-based prediction model and various other prompt engineering techniques, our method with GLM-4-Plus achieved the best performance on test dataset, attaining an F1-score of 0.89 and accuracy of 0.89. On the external validation dataset, F1-score (0.86) and accuracy (0.92) of the proposed method with GPT-4o were the highest. Compared to the popular strategy with manually selected typical samples (few-shot) and CoT designed by doctors (F1-score=0.83 and accuracy=0.83), the proposed method that summarized disease characteristics (F-Sum) based on LLM and automatically generated CoT performed better (F1-score=0.89 and accuracy=0.90). Although the BERT-based model got similar results on the test dataset (F1-score=0.85 and accuracy=0.88), its predictive performance significantly decreased on the external validation set (F1-score=0.48 and accuracy=0.78).
Conclusions: These findings highlight the potential of LLMs to revolutionize pulmonary disease prediction, particularly in resource-constrained settings, by surpassing traditional models in both accuracy and flexibility. The proposed prompt engineering strategy not only improves predictive performance but also enhances the adaptability of LLMs in complex medical contexts, offering a promising tool for advancing disease diagnosis and clinical decision-making.
Keywords: LLM; RAG; large language models; prompt engineering; pulmonary disease prediction; retrieval-augmented generation.
© Ronghao Li, Shuai Mao, Congmin Zhu, Yingliang Yang, Chunting Tan, Li Li, Xiangdong Mu, Honglei Liu, Yuqing Yang. Originally published in the Journal of Medical Internet Research (https://www.jmir.org).
Conflict of interest statement
Figures

Similar articles
-
Predicting 30-Day Postoperative Mortality and American Society of Anesthesiologists Physical Status Using Retrieval-Augmented Large Language Models: Development and Validation Study.J Med Internet Res. 2025 Jun 3;27:e75052. doi: 10.2196/75052. J Med Internet Res. 2025. PMID: 40460423 Free PMC article.
-
Developing an ICD-10 Coding Assistant: Pilot Study Using RoBERTa and GPT-4 for Term Extraction and Description-Based Code Selection.JMIR Form Res. 2025 Feb 11;9:e60095. doi: 10.2196/60095. JMIR Form Res. 2025. PMID: 39935026 Free PMC article.
-
Improving Large Language Models' Summarization Accuracy by Adding Highlights to Discharge Notes: Comparative Evaluation.JMIR Med Inform. 2025 Jul 24;13:e66476. doi: 10.2196/66476. JMIR Med Inform. 2025. PMID: 40705416 Free PMC article.
-
Examining the Role of Large Language Models in Orthopedics: Systematic Review.J Med Internet Res. 2024 Nov 15;26:e59607. doi: 10.2196/59607. J Med Internet Res. 2024. PMID: 39546795 Free PMC article.
-
Applications of Large Language Models in the Field of Suicide Prevention: Scoping Review.J Med Internet Res. 2025 Jan 23;27:e63126. doi: 10.2196/63126. J Med Internet Res. 2025. PMID: 39847414 Free PMC article.
References
-
- Daniel R, Jones H, Gregory JW, et al. Predicting type 1 diabetes in children using electronic health records in primary care in the UK: development and validation of a machine-learning algorithm. Lancet Digit Health. 2024 Jun;6(6):e386–e395. doi: 10.1016/S2589-7500(24)00050-5. doi. Medline. - DOI - PubMed
-
- Vaswani A, Shazeer N, Parmar N. Attention is all you need. arXiv. 2017 Jun 12; doi: 10.48550/arXiv.1706.03762. Preprint posted online on. doi. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

