Ventral hippocampus neurons encode meal-related memory
- PMID: 40500290
- PMCID: PMC12159138
- DOI: 10.1038/s41467-025-59687-1
Ventral hippocampus neurons encode meal-related memory
Abstract
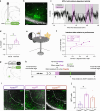
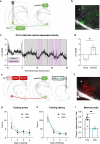
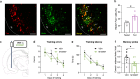
The ability to encode and retrieve meal-related information is critical to efficiently guide energy acquisition and consumption, yet the underlying neural processes remain elusive. Here we reveal that ventral hippocampus (HPCv) neuronal activity dynamically elevates between eating bouts during meal consumption and this response is predictive of performance in a foraging-related memory test for the spatial location of a previously consumed meal. Targeted recombination-mediated ablation of HPCv meal-responsive neurons impairs meal location memory without influencing food motivation or spatial memory for escape location. These HPCv meal-responsive neurons project to the lateral hypothalamic area (LHA) and are enriched in serotonin 2a receptors (5HT2aR). Either chemogenetic silencing of HPCv-to-LHA projections or intra-HPCv 5HT2aR antagonist yielded meal location memory deficits, as well as increased caloric intake driven by shorter temporal intervals between meals. Collective results identify a population of HPCv neurons in male rats that dynamically respond during eating to encode meal-related memories.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: B.C.R. and M.R.H. both receive research funding from Novo Nordisk and Boehringer Ingelheim that was not used in support of these studies. M.R.H. receives research funding from Eli Lilly & Co., Gila Therapeutics, and Pfizer that was not used in support of these studies. All other authors declare no competing interests.
Figures




Update of
-
Ventral hippocampus neurons encode meal-related memory.bioRxiv [Preprint]. 2023 Oct 10:2023.10.10.561731. doi: 10.1101/2023.10.10.561731. bioRxiv. 2023. Update in: Nat Commun. 2025 Jun 11;16(1):4898. doi: 10.1038/s41467-025-59687-1. PMID: 37873229 Free PMC article. Updated. Preprint.
References
-
- Higgs, S. Memory and its role in appetite regulation. Physiol. Behav.85, 67–72 (2005). - PubMed
-
- Higgs, S. & Donohoe, J. E. Focusing on food during lunch enhances lunch memory and decreases later snack intake. Appetite57, 202–206 (2011). - PubMed
-
- Collins, R. & Stafford, L. D. Feeling happy and thinking about food. Counteractive effects of mood and memory on food consumption. Appetite84, 107–112 (2015). - PubMed
MeSH terms
Grants and funding
- R01 DK104897/DK/NIDDK NIH HHS/United States
- R01 DK105155/DK/NIDDK NIH HHS/United States
- DK105155/U.S. Department of Health & Human Services | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (National Institute of Diabetes & Digestive & Kidney Diseases)
- DK104897/U.S. Department of Health & Human Services | NIH | National Institute of Diabetes and Digestive and Kidney Diseases (National Institute of Diabetes & Digestive & Kidney Diseases)
LinkOut - more resources
Full Text Sources
Medical

