Monitoring the rate and variability of somatic genomic alterations using long-read sequencing
- PMID: 40500302
- PMCID: PMC12159167
- DOI: 10.1038/s41598-025-01690-z
Monitoring the rate and variability of somatic genomic alterations using long-read sequencing
Abstract
Cancer initiation occurs when a cell acquires and accumulates mutations in genes involved in the regulation of cell processes: each cell division throughout a person's life introduces novel mutations in the cells' DNA and under normal circumstances, the body is primed to prevent those from leading to cancer. Occasionally, a subset of those mutations escapes those safeguards and might eventually result in the emergence of the disease. To understand the dynamics of accumulation of somatic mutations, we have performed longitudinal whole genome sequencing of DNA obtained from whole blood from healthy individuals and cancer patients using Oxford Nanopore Technologies' Long Read Sequencing. Here we show that the number of somatic single nucleotide variants detected increases with their age and that for specific mutational processes, changes can be detected within months. We computed aggregated metrics for unique participants at each timepoint across types of variants (based on single based substitution molecular signatures) and identified patterns of change both over an individual's lifespan (age) and over the sampling period (months). This study showcases the suitability of long read sequencing of blood DNA for detecting coarse-grained differences over time and enable future development of "state of the system" personalized prevention programs.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: All authors except Hag.L., D.A. and Han.L. are employees of the Ellison Institute, LLC. D.B.A. is a scientific advisor with equity interests in Oxford Nanopore Technologies. D.B.A. and N.M. are inventors on a provisional patent (US Provisional Patent Application no.: 63/581,553) related to this work. Oxford Nanopore Technologies contributed equipment, materials, reagents and technical support. The remaining authors declare no competing interests.
Figures


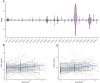
References
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials

