This is a preprint.
Synthetic essentiality of TRAIL/TNFSF10 in VHL-deficient renal cell carcinoma
- PMID: 40501957
- PMCID: PMC12154588
- DOI: 10.1101/2025.05.29.621197
Synthetic essentiality of TRAIL/TNFSF10 in VHL-deficient renal cell carcinoma
Abstract
Clear cell renal cell carcinoma (ccRCC) is the most common and aggressive subtype of kidney cancer. Loss of von Hippel-Lindau (VHL) and the consequent activation of hypoxia-inducible factor-α (HIFα, especially HIF2α) plays an essential role in ccRCC initiation and progression. The approved HIF2α inhibitor belzutifan faces the challenge of resistance, presenting an opportunity of co-targeting HIF2α and another vulnerability. This study elucidates the synthetic essentiality of TRAIL (tumor necrosis factor-related apoptosis-inducing ligand) in VHL-deficient ccRCC, uncovering a novel reciprocal regulation between HIF2α and TRAIL. TRAIL was identified as a direct transcriptional target of HIF2α and paradoxically found to be crucial for cell proliferation, primarily by activating the p38 MAPK pathway and facilitating G1/S phase transition. Depletion of endogenous TRAIL or inhibition of HIF2α with belzutifan sensitizes ccRCC cells to recombinant TRAIL, presenting a promising avenue for combination therapy to overcome both TRAIL resistance and belzutifan resistance in treating ccRCC.
Keywords: HIF2α; TRAIL; VHL; belzutifan; renal cell carcinoma; synthetic essentiality.
Conflict of interest statement
Competing interests: The authors declare no competing interests.
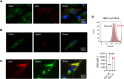
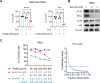
Figures




References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
