This is a preprint.
Decay drives RNA abundance regulation using three distinct regulatory mechanisms
- PMID: 40502020
- PMCID: PMC12157598
- DOI: 10.1101/2025.05.09.653099
Decay drives RNA abundance regulation using three distinct regulatory mechanisms
Abstract
RNA decay is essential for maintenance of normal RNA abundances; however how RNA decay is regulated to contribute to changes in RNA abundances is poorly understood. Here, we addressed this question by analyzing rates of RNA abundance change, RNA halflives (t 1/2s), and transcription rates in stimulated Arabidopsis leaf cells. This revealed three mechanisms by which decay influenced RNA abundance changes. First, the biggest changes in RNA abundances resulted from t 1/2 changes that reinforced transcriptional regulation (synergistic). Modest RNA abundance changes arose from a second mechanism in which t 1/2 changes opposed transcriptional regulation (oppositional). Finally, RNA decay alone also contributed to RNA abundance change, and RNA decay's measured capacity influence RNA abundances was similar to that of transcription. RNA decay also contributed to transcriptome homeostasis through stimulus-induced RNA buffering. Oppositional and buffering regulation shared key features, including excessive and commensurate rate changes, which suggested use of a shared regulatory mechanism which we call countercyclical regulation. In this study, countercyclical regulation was widespread and used for regulation of 90% of the RNAs with t 1/2 regulation.
Keywords: RNA buffering; RNA decay; RNA half-life (t1/2); countercyclical; nuclear-cytoplasmic coordination; transcription; transcriptome homeostasis; vascular transdifferentiation.
Figures


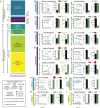


References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
