Multidrug-Resistant ESKAPEEc Pathogens From Bloodstream Infections in South Africa: A Cross-Sectional Study Assessing Resistance to WHO AWaRe Antibiotics
- PMID: 40510521
- PMCID: PMC12158662
- DOI: 10.1002/hsr2.70897
Multidrug-Resistant ESKAPEEc Pathogens From Bloodstream Infections in South Africa: A Cross-Sectional Study Assessing Resistance to WHO AWaRe Antibiotics
Abstract
Background and aims: Multidrug-resistant (MDR) pathogens, particularly members of the ESKAPE group and Escherichia coli (collectively referred to as ESKAPEEc), are major contributors to bloodstream infections (BSIs) and pose significant treatment challenges. This study aimed to characterize the antimicrobial resistance (AMR) profiles of ESKAPEEc isolates from BSIs in public hospitals in the uMgungundlovu District, South Africa, and to assess their resistance to World Health Organization (WHO) Access, Watch, and Reserve (AWaRe) antibiotics.
Methods: Between November 2017 and December 2018, blood samples (n = 195) were collected from adult and paediatric patients with suspected BSIs. Isolates were identified using the VITEK 2 system and confirmed by polymerase chain reaction (PCR). Antimicrobial susceptibility testing was performed using the Kirby-Bauer disk diffusion method and interpreted according to EUCAST/CLSI guidelines. The multiple antibiotic resistance index (MARI) was calculated. One-way analysis of variance (ANOVA) was used to assess associations between MARI and clinical variables, including ward type and facility level.
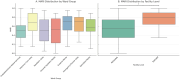
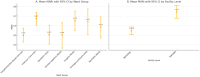
Results: Out of 195 presumptive isolates, 159 were confirmed as ESKAPEEc. The most frequently identified pathogens were Klebsiella pneumoniae (28.9%) and Staphylococcus aureus (28.3%). High resistance rates were observed across WHO Access and Watch antibiotics, including ampicillin (76% in E. coli), gentamicin (67.4% in K. pneumoniae), and ciprofloxacin (≥ 60% in most species). Carbapenem resistance in Acinetobacter baumannii reached 90%. Overall, 94.9% of isolates were MDR, and 93.1% had MARI ≥ 0.2. Significant differences in MARI values were observed across ward groups and facility levels, with the highest values recorded in intensive care units (mean = 0.67, 95% CI: 0.62-0.72) and tertiary hospitals (mean = 0.64, 95% CI: 0.60-0.68), compared to regional hospitals (mean = 0.52, 95% CI: 0.47-0.57).
Conclusion: The findings reveal a high burden of MDR ESKAPEEc in BSIs and widespread resistance to WHO Watch antibiotics. Targeted antimicrobial stewardship and the implementation of microbiology-guided therapy are urgently needed to optimize patient outcomes and curb the spread of resistance.
Keywords: MAR index; WHO AWaRe; antimicrobial resistance; bloodstream infections; multidrug resistance; pathogens.
© 2025 The Author(s). Health Science Reports published by Wiley Periodicals LLC.
Conflict of interest statement
S.Y.E. is a chairperson of the Global Respiratory Partnership and a member of the Global Hygiene Council, both supported by unrestricted educational grants from Reckitt (Pty.) Ltd. UK. The other authors declare no conflicts of interest.
Figures


Similar articles
-
Shifting trends and age distribution of ESKAPEEc resistance in bloodstream infection, Southwest China, 2012-2017.Antimicrob Resist Infect Control. 2019 Mar 29;8:61. doi: 10.1186/s13756-019-0499-1. eCollection 2019. Antimicrob Resist Infect Control. 2019. PMID: 30976388 Free PMC article.
-
Current trends in antimicrobial resistance of ESKAPEEc pathogens from bloodstream infections - Experience of a tertiary care centre in North India.Indian J Med Microbiol. 2024 Jul-Aug;50:100647. doi: 10.1016/j.ijmmb.2024.100647. Epub 2024 Jun 14. Indian J Med Microbiol. 2024. PMID: 38871082
-
Insights into bloodstream infections in South African paediatric burn patients: implications for antimicrobial stewardship.BMC Infect Dis. 2025 Mar 14;25(1):362. doi: 10.1186/s12879-025-10582-6. BMC Infect Dis. 2025. PMID: 40087547 Free PMC article.
-
Global Prevalence of Colistin Resistance in Klebsiella pneumoniae from Bloodstream Infection: A Systematic Review and Meta-Analysis.Pathogens. 2022 Sep 24;11(10):1092. doi: 10.3390/pathogens11101092. Pathogens. 2022. PMID: 36297149 Free PMC article. Review.
-
Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis.Lancet. 2022 Feb 12;399(10325):629-655. doi: 10.1016/S0140-6736(21)02724-0. Epub 2022 Jan 19. Lancet. 2022. PMID: 35065702 Free PMC article.
References
-
- WHO , “Bacterial Priority Pathogens List, Bacterial Pathogens of Public Health Importance to Guide Research, Development and Strategies to Prevent and Control Antimicrobial Resistance,” published 2024, https://www.who.int/publications/i/item/9789240093461. - PubMed
-
- De Angelis G., Fiori B., Menchinelli G., et al., “Incidence and Antimicrobial Resistance Trends in Bloodstream Infections Caused by ESKAPE and Escherichia coli at a Large Teaching Hospital in Rome, a 9‐Year Analysis (2007–2015),” European Journal of Clinical Microbiology & Infectious Diseases 37, no. 9 (2018): 1627–1636, 10.1007/s10096-018-3292-9. - DOI - PubMed
-
- Martinez R. M. and Wolk D. M., “Bloodstream Infections,” in Diagnostic Microbiology of the Immunocompromised Host, ed. Carroll K. C., Pfaller M. A., Landry M. L., McAdam A. J., Patel R., and Richter S. S., 1st ed. (ASM Press, 2016), 653–689, 10.1128/microbiolspec.DMIH2-0031-2016. - DOI
LinkOut - more resources
Full Text Sources

