Unraveling the interrelationship between breast cancer and endometriosis based on multi-omics analysis
- PMID: 40512411
- PMCID: PMC12165933
- DOI: 10.1007/s12672-025-02887-4
Unraveling the interrelationship between breast cancer and endometriosis based on multi-omics analysis
Abstract
Background: Endometriosis and breast cancer are significant global health burdens affecting women worldwide. Both conditions share notable characteristics including estrogen dependence, progressive growth patterns, recurrence tendencies, and metastatic potential. Despite these biological parallels, the molecular mechanisms connecting these conditions remain incompletely characterized. This study aimed to identify shared gene signatures and underlying molecular processes in breast cancer and endometriosis.
Methods: Expression matrices for both conditions were obtained from the Gene Expression Omnibus (GEO), UCSC Xena, and the Molecular Taxonomy of Breast Cancer International Consortium. Common differentially expressed genes (DEGs) were identified using the limma package. Comprehensive analyses included Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment, machine learning-based diagnostic and prognostic model development, potential therapeutic compound screening, tumor immune microenvironment (TIME) characterization, and hub gene identification with subsequent validation.
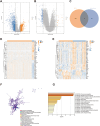
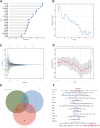
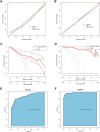
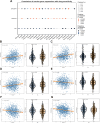
Results: The analysis identified 47 common DEGs between breast cancer and endometriosis. Functional assessment of these genes revealed their involvement in critical biological processes including cell cycle regulation, oxidative stress response, and secretory granule and recycling endosome dynamics. Integration of comprehensive genomic and clinical data led to the development of a prognostic model for breast cancer and a diagnostic model for endometriosis.
Conclusion: This study provides molecular insights into shared pathogenic mechanisms underlying breast cancer and endometriosis, highlighting common physiological pathways and key regulatory genes. These findings offer novel perspectives for understanding disease pathogenesis and potential therapeutic interventions for both conditions.
Keywords: Breast cancer; Endometriosis; Hub genes; Machine learning; Multi-omics analysis.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests. Ethical approval: Ethics approval not required. Consent to participate: Not applicable. Consent to publish: Not applicable.
Figures


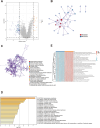

References
-
- Zondervan KT, Becker CM, Koga K, Missmer SA, Taylor RN, Viganò P. Endometriosis. Nat Rev Dis Primers. 2018. 10.1038/s41572-018-0008-5. - PubMed
-
- Taylor HS, Kotlyar AM, Flores VA. Endometriosis is a chronic systemic disease: clinical challenges and novel innovations. Lancet. 2021;397(10276):839–52. 10.1016/s0140-6736(21)00389-5. - PubMed
-
- Vercellini P, Viganò P, Somigliana E, Fedele L. Endometriosis: pathogenesis and treatment. Nat Reviews Endocrinol. 2013;10(5):261–75. 10.1038/nrendo.2013.255. - PubMed
LinkOut - more resources
Full Text Sources
