AbEpiTope-1.0: Improved antibody target prediction by use of AlphaFold and inverse folding
- PMID: 40512857
- PMCID: PMC12165000
- DOI: 10.1126/sciadv.adu1823
AbEpiTope-1.0: Improved antibody target prediction by use of AlphaFold and inverse folding
Abstract
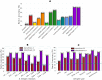
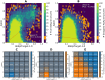
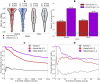
B cell epitope prediction tools are crucial for designing vaccines and disease diagnostics. However, predicting which antigens a specific antibody binds to and their exact binding sites (epitopes) remains challenging. Here, we present AbEpiTope-1.0, a tool for antibody-specific B cell epitope prediction, using AlphaFold for structural modeling and inverse folding for machine learning models. On a dataset of 1730 antibody-antigen complexes, AbEpiTope-1.0 outperforms AlphaFold in predicting modeled antibody-antigen interface accuracy. By creating swapped antibody-antigen complex structures for each antibody-antigen complex using incorrect antibodies, we show that predicted accuracies are sensitive to antibody input. Furthermore, a model variant optimized for antibody target prediction-differentiating true from swapped complexes-achieved an accuracy of 61.21% in correctly identifying antibody-antigen pairs. The tool evaluates hundreds of structures in minutes, providing researchers with a resource for screening antibodies targeting specific antigens. AbEpiTope-1.0 is freely available as a web server and software.
Figures


References
-
- Kozlova E. E. G., Cerf L., Schneider F. S., Viart B. T., Nguyen C., Steiner B. T., de Almeida Lima S., Molina F., Duarte C. G., Felicori L., Chávez-Olórtegui C., Machado-de-Ávila R. A., Computational B-cell epitope identification and production of neutralizing murine antibodies against Atroxlysin-I. Sci. Rep. 8, 14904 (2018). - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

