Antarctic ice-free terrestrial microbial functional redundancy in core ecological functions and microhabitat-specific microbial taxa and adaptive strategy
- PMID: 40514722
- PMCID: PMC12166587
- DOI: 10.1186/s40793-025-00735-0
Antarctic ice-free terrestrial microbial functional redundancy in core ecological functions and microhabitat-specific microbial taxa and adaptive strategy
Abstract
Background: Although ice-free terrestrial ecosystems in Antarctica cover only limited areas, they harbor diverse and metabolically active microbial communities. These ecosystems encompass distinct microhabitats such as mosses, lichens, and soils, each offering unique ecological niches. However, how different microbial taxa respond to microhabitat heterogeneity, ecological strategies such as functional redundancy and specialization contribute to adaptation in extreme environments remains underexplored. To address these questions, we employed high-throughput 16 S rRNA gene and ITS sequencing in combination with GeoChip-based functional gene profiling to assess the structure and functional potential of microbial communities across moss, lichen and soil microhabitats in Antarctic ice-free terrestrial ecosystem.
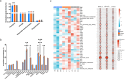
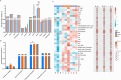
Results: Microhabitat type has a greater influence on microbial community structure and function in the ice-free Antarctic terrestrial ecosystem than geographical location. Though all prokaryotic communities were dominated by Pseudomonadota, Nostoc and Endobacter were significantly enriched in the moss and lichen microhabitats, respectively. Meanwhile, all fungal communities were primarily dominated by Ascomycota and Basidiomycota, with Byssoloma and Usnea showing significant enrichment in the moss and lichen microhabitats, respectively. Despite these taxonomic differences, the three microhabitats show similar core ecological functions with widespread microbial functional redundancy. Nevertheless, clear microhabitat-specific functional specialization was suggested. For example, moss microhabitat was enriched in Pyoverdin_pvcC and Zeaxanthin_glucosyltransferase, sdhA, lichen microhabitat harbored higher levels of nhaA, nikC, vacuolar_iron_transport, mttB, glucoamylase, pel_Cdeg, pme_Cdeg, rgh, rgl, while soil microhabitat was enriched in 5f1_ppn and isopullulanase. Notably, genes involved in carotenoid biosynthesis were significantly more abundant in moss and lichen microhabitats than in soil microhabitat, indicating the adaptive capacity of symbiotic microorganisms to mitigate ultraviolet radiation and oxidative stress to protect their hosts.
Conclusions: Microbial communities associated with distinct microhabitats (i.e. mosses, lichens, and soils) in Antarctic ice-free terrestrial ecosystem exhibit both functional redundancy in core ecological functions and microhabitat-specific specialization in key microbial taxa and adaptive strategy.
Keywords: Geochip functional microarray; High-throughput sequencing; Ice-free Antarctic terrestrial ecosystem; Lichen; Microbiome; Moss; Soil.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures





Similar articles
-
Species-specific effects of passive warming in an Antarctic moss system.R Soc Open Sci. 2019 Nov 13;6(11):190744. doi: 10.1098/rsos.190744. eCollection 2019 Nov. R Soc Open Sci. 2019. PMID: 31827828 Free PMC article.
-
Distinct microbial communities under different rock-associated microhabitats in the Qaidam Desert.Environ Res. 2024 Jun 1;250:118462. doi: 10.1016/j.envres.2024.118462. Epub 2024 Feb 15. Environ Res. 2024. PMID: 38367835
-
Adaptation to Environmental Extremes Structures Functional Traits in Biological Soil Crust and Hypolithic Microbial Communities.mSystems. 2022 Aug 30;7(4):e0141921. doi: 10.1128/msystems.01419-21. Epub 2022 Jul 19. mSystems. 2022. PMID: 35852333 Free PMC article.
-
Ecological Characteristics of Antarctic Fungi.Dokl Biol Sci. 2023 Feb;508(1):32-54. doi: 10.1134/S0012496622700120. Epub 2023 Apr 25. Dokl Biol Sci. 2023. PMID: 37186046 Review.
-
Emerging biological archives can reveal ecological and climatic change in Antarctica.Glob Chang Biol. 2022 Nov;28(22):6483-6508. doi: 10.1111/gcb.16356. Epub 2022 Sep 8. Glob Chang Biol. 2022. PMID: 35900301 Free PMC article. Review.
References
-
- Cary SC, McDonald IR, Barrett JE, Cowan DA. On the rocks: the microbiology of Antarctic dry Valley soils. Nat Rev Microbiol. 2010;8:129–38. - PubMed
-
- Chown SL, Clarke A, Fraser CI, Cary SC, Moon KL, McGeoch MA. The changing form of Antarctic biodiversity. Nature. 2015;522:431–8. - PubMed
-
- Camara P, Convey P, Rangel SB, Konrath M, Barreto CC, Pinto OHB, et al. The largest moss carpet transplant in Antarctica and its bryosphere cryptic biodiversity. Extremophiles. 2021;25:369–84. - PubMed
-
- Grube M, Cardinale M, de Castro JV Jr., Muller H, Berg G. Species-specific structural and functional diversity of bacterial communities in lichen symbioses. ISME J. 2009;3:1105–15. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
