Methanol-Free Protein Expression in Komagataella phaffii With Magnetic or Non-Magnetic Heating
- PMID: 40515638
- PMCID: PMC12166525
- DOI: 10.1111/1751-7915.70183
Methanol-Free Protein Expression in Komagataella phaffii With Magnetic or Non-Magnetic Heating
Abstract
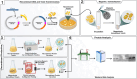
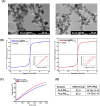
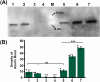
Komagataella phaffii is among the most widely used expression systems, with methanol-inducible promoters being preferred for protein expression due to their stringent regulation and exceptional strength. However, the applicability of this system, particularly in food and pharmaceutical products, is limited by methanol's toxic and pro-inflammatory properties. Therefore, obtaining a novel methanol-free expression system is necessary. In this study, we obtained a novel expression plasmid, pHSPαA, carrying the HSP70 promoter (PHSP70) to regulate heterologous expression through heat induction. The extracellular expression of azurin was achieved using this methanol-free system under the control of PHSP70, induced by either magnetic or non-magnetic heating. To enhance heat-induced expression, recombinant cells were immobilised with Fe3O4@PEI25 kDa nanoparticles, which facilitated heat release under an AC magnetic field, thereby increasing cell permeability and protein secretion. A time-dependent increase in protein expression was observed in non-magnetic heating but not under magnetic heating. However, immobilised cells exhibited a higher protein secretion capacity compared to non-immobilised cells. These findings suggest that the novel methanol-free expression system represents a promising alternative for heterologous gene expression, particularly for the production of therapeutically relevant and food-grade recombinant proteins.
Keywords: Komagataella phaffii; HSP70 promoter; alternating magnetic field; magnetic nanoparticles; methanol‐free expression.
© 2025 The Author(s). Microbial Biotechnology published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Bi-directionalized promoter systems allow methanol-free production of hard-to-express peroxygenases with Komagataella Phaffii.Microb Cell Fact. 2024 Jun 15;23(1):177. doi: 10.1186/s12934-024-02451-9. Microb Cell Fact. 2024. PMID: 38879507 Free PMC article.
-
Knock-out of the major regulator Flo8 in Komagataella phaffii results in unique host strain performance for methanol-free recombinant protein production.N Biotechnol. 2024 Dec 25;84:105-114. doi: 10.1016/j.nbt.2024.10.001. Epub 2024 Oct 9. N Biotechnol. 2024. PMID: 39384085
-
Scalable protein production by Komagataella phaffii enabled by ARS plasmids and carbon source-based selection.Microb Cell Fact. 2024 Apr 20;23(1):116. doi: 10.1186/s12934-024-02368-3. Microb Cell Fact. 2024. PMID: 38643119 Free PMC article.
-
Recent progress on heterologous protein production in methylotrophic yeast systems.World J Microbiol Biotechnol. 2024 May 11;40(7):200. doi: 10.1007/s11274-024-04008-9. World J Microbiol Biotechnol. 2024. PMID: 38730212 Free PMC article. Review.
-
Advancing recombinant protein expression in Komagataella phaffii: opportunities and challenges.FEMS Yeast Res. 2025 Jan 30;25:foaf010. doi: 10.1093/femsyr/foaf010. FEMS Yeast Res. 2025. PMID: 40074550 Free PMC article. Review.
References
-
- Acar, M. , Abul N., Yildiz S., et al. 2022. “Affinity‐Based and in a Single Step Purification of Recombinant Horseradish Peroxidase A2A Isoenzyme Produced by Pichia pastoris .” Bioprocess and Biosystems Engineering 46: 523–534. - PubMed
-
- Ádám, A. , Bártfai R., Lele Z., Krone P. H., and Orbán L.. 2000. “Heat‐Inducible Expression of a Reporter Gene Detected by Transient Assay in Zebrafish.” Experimental Cell Research 256: 282–290. - PubMed
-
- Ansari, F. , Grigoriev P., Libor S., Tothill I. E., and Ramsden J. J.. 2009. “DBT Degradation Enhancement by Decorating Rhodococcus erythropolis IGST8 With Magnetic Fe3O4 Nanoparticles.” Biotechnology and Bioengineering 102: 1505–1512. - PubMed
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

