Comparative Genome Analysis of Three Halobacillus Strains Isolated From Saline Environments Reveal Potential Salt Tolerance and Algicidal Mechanisms
- PMID: 40518659
- PMCID: PMC12167742
- DOI: 10.1111/1758-2229.70121
Comparative Genome Analysis of Three Halobacillus Strains Isolated From Saline Environments Reveal Potential Salt Tolerance and Algicidal Mechanisms
Abstract
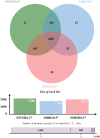
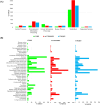
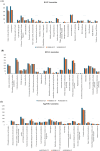
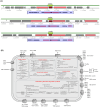
Harmful algal blooms (HABs) pose a significant global threat to water ecosystems, prompting extensive research into their inhibition and control strategies. This study presents genomic and bioinformatic analyses to investigate the algicidal potential and elucidate the survival mechanisms in harsh conditions of newly identified Halobacillus species three strains (SSTM10-2T, SSBR10-3T, and SSHM10-5T) isolated from saline environments. Moreover, genomic and bioinformatic analyses were conducted to elucidate their survival mechanisms in harsh conditions. Moreover, comparative genomic analysis revealed a diverse set of orthologous genes, with a core genome primarily associated with metabolism and information processing. Pangenome analysis highlighted accessory and unique genes potentially involved in environmental adaptation and stress response. Functional annotation using KEGG pathways identified genes linked to xenobiotic compound degradation, stress tolerance, and salt adaptation. Additionally, the study elucidated potential mechanisms underlying algicidal activity, implicating Carbohydrate-Active enZYmes (CAZymes), cytochrome P450 oxidases (CYP), and quorum sensing (QS) systems. Finally, analysis of KEGG pathways related to microcystin degradation suggested the strains' capacity to mitigate HABs. Thus, this research enhances understanding of the genomic diversity, phylogeny, and functional characteristics of Halobacillus species, offering insights into their ecological roles and potential applications in biotechnology and environmental management.
Keywords: Halobacillus sp.; algicidal activity; comparative genome analysis; halotolerance; stress response.
© 2025 The Author(s). Environmental Microbiology Reports published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


Similar articles
-
Halobacillus salinarum sp. nov., Halobacillus shinanisalinarum sp. nov. and Halobacillus amylolyticus sp. nov., isolated from saltern soil.Int J Syst Evol Microbiol. 2023 Oct;73(10). doi: 10.1099/ijsem.0.006098. Int J Syst Evol Microbiol. 2023. PMID: 37882660
-
Salicibibacter halophilus sp. nov., a moderately halophilic bacterium isolated from kimchi.J Microbiol. 2019 Nov;57(11):997-1002. doi: 10.1007/s12275-019-9421-z. Epub 2019 Oct 28. J Microbiol. 2019. PMID: 31659686
-
Halobacillus hunanensis sp. nov., a moderately halophilic bacterium isolated from a subterranean brine.Antonie Van Leeuwenhoek. 2009 Nov;96(4):497-504. doi: 10.1007/s10482-009-9365-0. Epub 2009 Jul 18. Antonie Van Leeuwenhoek. 2009. PMID: 19618289
-
Diverse interactions between bacteria and microalgae: A review for enhancing harmful algal bloom mitigation and biomass processing efficiency.Heliyon. 2024 Aug 24;10(17):e36503. doi: 10.1016/j.heliyon.2024.e36503. eCollection 2024 Sep 15. Heliyon. 2024. PMID: 39286093 Free PMC article. Review.
-
Unveiling the trifecta of cyanobacterial quorum sensing: LuxI, LuxR and LuxS as the intricate machinery for harmful algal bloom formation in freshwater ecosystems.Sci Total Environ. 2024 May 10;924:171644. doi: 10.1016/j.scitotenv.2024.171644. Epub 2024 Mar 11. Sci Total Environ. 2024. PMID: 38471587 Review.
References
-
- Agamennone, V. , Le N. G., van Straalen N. M., Brouwer A., and Roelofs D.. 2019. “Antimicrobial Activity and Carbohydrate Metabolism in the Bacterial Metagenome of the Soil‐Living Invertebrate <styled-content style="fixed-case"> Folsomia candida </styled-content> .” Scientific Reports 9: 7308. 10.1038/s41598-019-43828-w. - DOI - PMC - PubMed
-
- An, S. Y. , Asahara M., Goto K., Kasai H., and Yokota A.. 2007. “ <styled-content style="fixed-case"> Terribacillus saccharophilus </styled-content> Gen. Nov., sp. Nov. and <styled-content style="fixed-case"> Terribacillus halophilus </styled-content> sp. Nov., Spore‐Forming Bacteria Isolated From Field Soil in Japan.” International Journal of Systematic and Evolutionary Microbiology 57: 51–55. 10.1099/ijs.0.64340-0. - DOI - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

