Potential cerebrospinal fluid metabolomic biomarkers and early prediction model for Parkinson's disease
- PMID: 40520532
- PMCID: PMC12163037
- DOI: 10.3389/fnagi.2025.1582362
Potential cerebrospinal fluid metabolomic biomarkers and early prediction model for Parkinson's disease
Abstract
Objective: To identify key cerebrospinal fluid (CSF) metabolomic biomarkers associated with Parkinson's disease (PD) and prodromal PD, providing insights for intervention strategy development.
Methods: Six hundred and thirty-nine participants from the Parkinson's Progression Markers Initiative (PPMI) cohort were included: 300 PD patients, 112 healthy controls (HC), and 227 prodromal PD patients. Regression analysis and OPLS-DA identified metabolic biomarkers, while pathway analysis examined their relationship to clinical features. An XGBoost-based early prediction model was developed to assess the distinction between PD, prodromal PD, and HC. A two-sample bidirectional Mendelian randomization analysis was performed to examine the association between differential metabolites and Parkinson's disease.
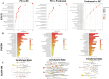
Results: Sixty-four metabolites were significantly altered in PD patients compared to HC, with 58 elevated and 6 reduced. In prodromal PD, 19 metabolites were increased, and 34 were decreased. Key metabolic pathways involved glutathione and amino acid metabolism. Dopamine 3-O-sulfate correlated with PD progression, levodopa-equivalent dose, and non-motor symptoms. The XGBoost model demonstrated high specificity in predicting the onset of PD. The MR analysis results showed a positive correlation between higher genetic predictions of dopamine 3-O-sulfate levels and the risk of Parkinson's disease. In contrast, the reverse MR analysis found that Parkinson's disease susceptibility significantly increased dopamine 3-O-sulfate levels.
Conclusion: The differential expression of CSF metabolites reveals early cellular metabolic changes, providing insights for early diagnosis and monitoring PD progression. A bidirectional causal relationship exists between genetically determined PD susceptibility and metabolites.
Keywords: PD susceptibility; Parkinson diseases; bidirectional Mendelian randomization; early prediction model; metabolomic biomarkers.
Copyright © 2025 Zhang, Yan, Kong, Zhang and Su.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Similar articles
-
Causal association between cerebrospinal fluid metabolites and Parkinson's disease: A two-sample bidirectional mendelian randomization study.Behav Brain Res. 2025 Mar 28;482:115426. doi: 10.1016/j.bbr.2025.115426. Epub 2025 Jan 8. Behav Brain Res. 2025. PMID: 39793738
-
CSF biomarkers associated with disease heterogeneity in early Parkinson's disease: the Parkinson's Progression Markers Initiative study.Acta Neuropathol. 2016 Jun;131(6):935-49. doi: 10.1007/s00401-016-1552-2. Epub 2016 Mar 28. Acta Neuropathol. 2016. PMID: 27021906 Free PMC article.
-
Longitudinal analyses of cerebrospinal fluid α-Synuclein in prodromal and early Parkinson's disease.Mov Disord. 2019 Sep;34(9):1354-1364. doi: 10.1002/mds.27806. Epub 2019 Jul 30. Mov Disord. 2019. PMID: 31361367 Free PMC article.
-
Cerebrospinal Fluid Amyloid β1-42, Tau, and Alpha-Synuclein Predict the Heterogeneous Progression of Cognitive Dysfunction in Parkinson's Disease.J Mov Disord. 2016 May;9(2):89-96. doi: 10.14802/jmd.16017. Epub 2016 May 25. J Mov Disord. 2016. PMID: 27240810 Free PMC article. Review.
-
Approaches to Early Parkinson's Disease Subtyping.J Parkinsons Dis. 2024;14(s2):S297-S306. doi: 10.3233/JPD-230419. J Parkinsons Dis. 2024. PMID: 39331104 Free PMC article. Review.
References
LinkOut - more resources
Full Text Sources

