Construction of epilepsy diagnosis model based on cell senescence-related genes and its potential mechanism
- PMID: 40520613
- PMCID: PMC12162300
- DOI: 10.3389/fneur.2025.1555586
Construction of epilepsy diagnosis model based on cell senescence-related genes and its potential mechanism
Abstract
Introduction: Epilepsy is a chronic brain disease with a certain degree of neurodegeneration and is caused by abnormal discharges of neurons. The mechanism of cell senescence has garnered increasing attention in neurodegenerative diseases. However, the role of cell senescence in the onset and progression of epilepsy is unclear. Therefore, this study constructed a diagnostic model of epilepsy based on cellular senescence-related genes (CSRGs) to analyze their role in disease pathogenesis.
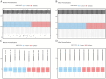
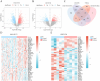
Methods: The differentially expressed genes (DEGs) were screened from the epileptic sample dataset of the gene expression omnibus (GEO) database, and the cellular senescence-related DEGs (CSRDEGs) related to epilepsy were identified by CSRGs crossover. The functional enrichment characteristics of CSRDEGs were analyzed using gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG) enrichment analyses. The differences in biological processes between high and low-risk groups were analyzed using gene set enrichment analysis (GSEA). For model construction, logistic regression, random forest, and least absolute shrinkage and selection operator (LASSO) regression were employed to identify key genes, including ribosomal protein S6 kinase alpha-3 (RPS6KA3), cathepsin D (CTSD), and zinc finger protein 101 (ZNF101). Subsequently, a multifactor logistic regression model was developed to evaluate the risk of epilepsy based on these screened genes.
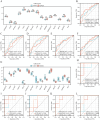
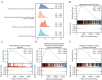
Results: The model exhibited higher area under the curve (AUC) values in the GSE data sets 143272 and 32534, producing encouraging results. Finally, mRNA-miRNA and mRNA-transcription factors (TFs) networks revealed the potential regulatory mechanism of the selected critical genes in the disease.
Discussion: This study elucidated the possible process of cell senescence in epileptic diseases through bioinformatics analysis, offering a potential target for personalized diagnosis and precise treatment.
Keywords: bioinformatics analysis; cell senescence; diagnosis model; epilepsy; senescence-associated secretory phenotype.
Copyright © 2025 Gong, Lu, Xiao, Wang, Cui and Tang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





Similar articles
-
Identification and validation of senescence-related genes in circulating endothelial cells of patients with acute myocardial infarction.Front Cardiovasc Med. 2022 Dec 13;9:1057985. doi: 10.3389/fcvm.2022.1057985. eCollection 2022. Front Cardiovasc Med. 2022. PMID: 36582740 Free PMC article.
-
Exosome-related gene identification and diagnostic model construction in hepatic ischemia-reperfusion injury.Sci Rep. 2024 Sep 28;14(1):22450. doi: 10.1038/s41598-024-73441-5. Sci Rep. 2024. PMID: 39341981 Free PMC article.
-
[Exploration of key ferroptosis-related genes as therapeutic targets for sepsis based on bioinformatics and the depiction of their immune profiles characterization].Zhonghua Wei Zhong Bing Ji Jiu Yi Xue. 2024 Oct;36(10):1025-1032. doi: 10.3760/cma.j.cn121430-20240524-00457. Zhonghua Wei Zhong Bing Ji Jiu Yi Xue. 2024. PMID: 39586719 Chinese.
-
A senescence-based prognostic gene signature for colorectal cancer and identification of the role of SPP1-positive macrophages in tumor senescence.Front Immunol. 2023 Apr 6;14:1175490. doi: 10.3389/fimmu.2023.1175490. eCollection 2023. Front Immunol. 2023. PMID: 37090726 Free PMC article.
-
The Significance of Cellular Senescence Hub Genes in the Diagnosis and Subtype Classification of a Comprehensive Database of Gene Expression in Intervertebral Disc Degeneration.JOR Spine. 2025 Mar 6;8(1):e70050. doi: 10.1002/jsp2.70050. eCollection 2025 Mar. JOR Spine. 2025. PMID: 40061887 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

