A survey of computational approaches for characterizing microbial interactions in microbial mats
- PMID: 40524188
- PMCID: PMC12168310
- DOI: 10.1186/s13059-025-03634-2
A survey of computational approaches for characterizing microbial interactions in microbial mats
Abstract
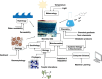
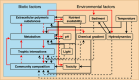
In this review, we use microbial mat communities as a general model system to highlight the strengths and limitations of current computational methods for analyzing interactions between members of microbial ecosystems. We describe the factors that make this environment have such a high degree of interaction, and we explore different categories of both laboratory and computational tools for studying these interactions. For each tool, we describe efforts to apply them to microbial mats in the past and, in the process, argue that genome-scale metabolic models have breakthrough potential for modeling microbial interactions in microbial mats.
Keywords: Comparative genomics; Computational models; Environmental data; Meta-omics; Metabolic models; Microbial ecology; Microbial evolution; Microbial mats; Transdisciplinary studies.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare that they have no competing interests.
Figures

Similar articles
-
Determinism and stochasticity drive microbial community assembly and microbial interactions in calcareous glacier forefields.Appl Environ Microbiol. 2025 Jun 18;91(6):e0030225. doi: 10.1128/aem.00302-25. Epub 2025 May 15. Appl Environ Microbiol. 2025. PMID: 40372053 Free PMC article.
-
Assessing the comparative effects of interventions in COPD: a tutorial on network meta-analysis for clinicians.Respir Res. 2024 Dec 21;25(1):438. doi: 10.1186/s12931-024-03056-x. Respir Res. 2024. PMID: 39709425 Free PMC article. Review.
-
Integrating Gut Microbiome and Metabolomics with Magnetic Resonance Enterography to Advance Bowel Damage Prediction in Crohn's Disease.J Inflamm Res. 2025 Jun 11;18:7631-7649. doi: 10.2147/JIR.S524671. eCollection 2025. J Inflamm Res. 2025. PMID: 40535353 Free PMC article.
-
A bloom of a single bacterium shapes the microbiome during outdoor diatom cultivation collapse.mSystems. 2025 Jun 17;10(6):e0037525. doi: 10.1128/msystems.00375-25. Epub 2025 May 14. mSystems. 2025. PMID: 40366134 Free PMC article.
-
Radiation-induced injury and the gut microbiota: insights from a microbial perspective.Therap Adv Gastroenterol. 2025 Jun 16;18:17562848251347347. doi: 10.1177/17562848251347347. eCollection 2025. Therap Adv Gastroenterol. 2025. PMID: 40535532 Free PMC article. Review.
References
-
- Jean-Pierre F, Henson MA, O’Toole GA. Metabolic modeling to interrogate microbial disease: a tale for experimentalists. Front Mol Biosci. 2021;8. Available from: https://www.frontiersin.org/articles/10.3389/fmolb.2021.634479. Cited 2022 Aug 25. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

