The RNF/NQR redox pumps: a versatile system for energy transduction in bacteria and archaea
- PMID: 40528048
- PMCID: PMC12174285
- DOI: 10.1007/s00253-025-13531-0
The RNF/NQR redox pumps: a versatile system for energy transduction in bacteria and archaea
Abstract
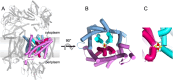
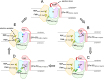
The Na+ (or H+)-translocating ferredoxin:NAD+ oxidoreductase (also called RNF, rhodobacter nitrogen fixation, complex) catalyzes the oxidation of reduced ferredoxin with NAD+, hereby generating an electrochemical gradient. In the reverse reaction driven by an electrochemical gradient, RNF provides reduced ferredoxin using NADH as electron donor. RNF plays a crucial role in the metabolism of many anaerobes, such as amino acid fermenters, acetogens, or aceticlastic methanogens. The Na+-translocating NADH:quinone oxidoreductase (NQR), which has evolved from an RNF, is found in selected bacterial groups including anaerobic, marine, or pathogenic organisms. Since NQR and RNF are not related to eukaryotic respiratory complex I (NADH:quinone oxidoreductase), members of this oxidoreductase family are promising targets for novel antibiotics. RNF and NQR share a membrane-bound core complex consisting of four subunits, which represent an essential functional module for redox-driven cation transport. Several recent 3D structures of RNF and NQR in different states put forward conformational coupling of electron transfer and Na+ translocation reaction steps. Based on this common principle, putative reaction mechanisms of RNF and NQR redox pumps are compared. KEY POINTS: • Electrogenic ferredoxin:NAD+ oxidoreductases (RNF complexes) are found in bacteria and archaea. • The Na+ -translocating NADH:quinone oxidoreductase (NQR) is evolutionary related to RNF. • The mechanism of energy conversion by RNF/NQR complexes is based on conformational coupling of electron transfer and cation transport reactions.
Keywords: Electrochemical proton gradient; Electrochemical sodium gradient; Electron transport; NQR; RNF; Respiration.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethical approval: Not applicable. Competing interests: The authors declare no competing interests.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

