Protocol for spatial proteomic profiling of tonsil cancer microenvironments using multiplexed imaging-powered deep visual proteomics
- PMID: 40536875
- PMCID: PMC12221282
- DOI: 10.1016/j.xpro.2025.103901
Protocol for spatial proteomic profiling of tonsil cancer microenvironments using multiplexed imaging-powered deep visual proteomics
Abstract
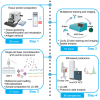
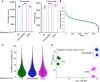
Here, we present a protocol for spatial proteomic profiling of the tumor microenvironment in tonsil cancer using multiplexed imaging-powered deep visual proteomics (mipDVP). We describe steps for automated 22-plex immunofluorescence staining and imaging on formalin-fixed paraffin-embedded (FFPE) tissue sections, automated single-cell laser microdissection, and single-cell-type mass spectrometry. This workflow enables the spatially resolved isolation of distinct cell populations for proteomic analysis. We optimized this protocol for studying tumor-immune interactions, where it facilitates the systematic identification of biomarkers and functional cellular networks. For complete details on the use and execution of this protocol, please refer to Zheng et al.1.
Keywords: Biotechnology and bioengineering; Cancer; Microscopy.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests M.M. is an indirect investor in Evosep Biosystems and OmicVision Biosciences. A.M. is a co-founder and chief scientific officer of OmicVision Biosciences.
Figures



References
MeSH terms
LinkOut - more resources
Full Text Sources

