MVHGCN: Predicting circRNA-disease associations with multi-view heterogeneous graph convolutional neural networks
- PMID: 40536898
- PMCID: PMC12225982
- DOI: 10.1371/journal.pcbi.1013225
MVHGCN: Predicting circRNA-disease associations with multi-view heterogeneous graph convolutional neural networks
Abstract
Circular RNA, a class of RNA molecules gaining widespread attentions, has been widely recognized as a potential biomarker for many diseases. In recent years, significant progress has been made in the study of the associations between circRNA and diseases. However, traditional experimental methods are often inefficient and costly, making computational models an effective alternative. Nevertheless, existing computational methods still face challenges such as data sparsity and the difficulty of confirming negative samples, which limits the accuracy of predictions. To address these challenges, a novel computational method, namely MVHGCN, is proposed based on multi-view and graph convolutional networks to predict potential associations between circRNA and diseases. MVHGCN first constructs a heterogeneous graph and generates feature descriptors by integrating multiple databases. Then it extracts different connection views of circRNA and diseases through meta-paths, maximizing the utilization of known association information, and aggregates deep feature information through graph convolutional networks. Finally, a MLP is used to predict the association scores. The experimental results show that MVHGCN significantly outperforms existing methods on benchmark datasets by 5-fold cross-validation. This research provides an effective new approach to studying the associations between circRNAs and diseases, capable of alleviating the problem of data sparsity and accurately identifying potential associations.
Copyright: © 2025 Miao et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures


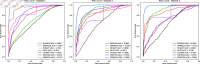
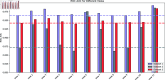
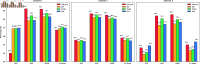
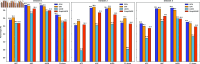
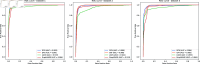

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

