Genomic evidence for flies as carriers of zoonotic pathogens on dairy farms
- PMID: 40537478
- PMCID: PMC12179284
- DOI: 10.1038/s41522-025-00685-y
Genomic evidence for flies as carriers of zoonotic pathogens on dairy farms
Abstract
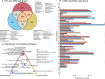
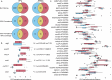
Dairy farms are major reservoirs of zoonotic bacterial pathogens, which harbor antimicrobial resistance genes (ARGs), and raise critical questions about their dissemination on and off the farm environment. Here, we investigated the role of coprophagous muscid flies (Diptera: Muscidae) as carriers of zoonotic pathogens and antimicrobial resistance. We collected cow manure and flies on a dairy farm and used shotgun metagenomics to identify the presence of clinically relevant bacteria, virulence factors, and ARGs in both environments. Our results reveal that, although the fly microbiome is largely composed of manure-associated taxa, they also harbor specific insect-associated bacteria, which may be involved in nutrient provisioning to the host. Furthermore, we identifed shared ARGs, virulence factors, and zoonotic pathogens enriched within the fly gastrointestinal tract (GIT). Our study illustrates the potential flow of pathogenic microorganisms from manure to coprophagous flies, suggesting that flies may pose an important zoonotic threat on dairy farms.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Ethical approval: This study exclusively involved coprophagous muscid flies (Diptera: Muscidae) and did not involve any vertebrates or higher invertebrates. All procedures, including anesthesia and dissection, were conducted in accordance with standard ethical guidelines for invertebrate research. As Muscidaeare not classified as higher invertebrates or regulated animals under institutional or national guidelines, no formal ethical approval was required.
Figures
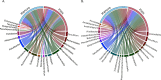

Similar articles
-
Phylogenetic intermixing reveals stable fly-mediated circulation of mastitis-associated bacteria in dairy settings.mSystems. 2025 Aug 1:e0021525. doi: 10.1128/msystems.00215-25. Online ahead of print. mSystems. 2025. PMID: 40748061
-
Potential pathogens and antimicrobial resistance genes in household environments: a study of soil floors and cow dung in rural Bangladesh.Appl Environ Microbiol. 2025 Jun 18;91(6):e0066925. doi: 10.1128/aem.00669-25. Epub 2025 May 27. Appl Environ Microbiol. 2025. PMID: 40422288 Free PMC article.
-
Bacterial abundance and antimicrobial resistance prevalence carried by adult house flies (Diptera: Muscidae) at Kansas dairy and beef cattle operations.J Med Entomol. 2025 Jul 17;62(4):984-994. doi: 10.1093/jme/tjaf052. J Med Entomol. 2025. PMID: 40261132
-
Prevalence of zoonotic or potentially zoonotic bacteria, antimicrobial resistance, and somatic cell counts in organic dairy production: current knowledge and research gaps.Foodborne Pathog Dis. 2009 Jun;6(5):525-39. doi: 10.1089/fpd.2008.0181. Foodborne Pathog Dis. 2009. PMID: 19422303
-
Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19.Cochrane Database Syst Rev. 2022 May 20;5(5):CD013665. doi: 10.1002/14651858.CD013665.pub3. Cochrane Database Syst Rev. 2022. PMID: 35593186 Free PMC article.
References
-
- Call, D. R., Davis, M. A. & Sawant, A. A. Antimicrobial resistance in beef and dairy cattle production. Anim. Health Res. Rev.9, 159–167 (2008). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

