Detection assay of polymyxin resistance coding mcr-1 gene based on CRISPR/Cas13a system
- PMID: 40538650
- PMCID: PMC12176852
- DOI: 10.3389/fcimb.2025.1553681
Detection assay of polymyxin resistance coding mcr-1 gene based on CRISPR/Cas13a system
Abstract
Introduction: Polymyxins are reserved as an ultimate defense against multidrug-resistant bacteria. The emergence of the polymyxin resistance gene mcr-1 poses a potential risk for the treatment of severe infections caused by Gram-negative bacteria. Timely detection and monitoring the mcr-1 gene are essential for guiding anti-infective therapy and controlling the spread of polymyxin resistance. Quantitative real-time PCR (qPCR) is one of the common methods for detecting resistance genes. However, qPCR has equipment dependency, and is not feasible in primary healthcare settings. Currently, there remains a lack of a highly sensitive and portable method for detecting the mcr-1 gene.
Methods: We established and optimized detection assays of the mcr-1 gene based on CRISPR/Cas13a system and lateral flow strips. The detection method was preliminarily evaluated using clinical isolates from Escherichia coli, compared with qPCR.
Results: The method for detecting the mcr-1 gene based on the CRISPR/Cas13a system and lateral flow strips was established, with a detection limit of 100 copies/mL. This method demonstrated high analytical specificity, with no cross-reactivity detected in non-mcr-1 and non-resistant strains. Among 36 clinical isolates, the method identified 31 strains as positive for the mcr-1 gene, and had a 100% concordance rate with the results of qPCR.
Conclusions: We established a detection method for the polymyxin resistance mcr-1 gene based on the CRISPR/Cas13a system. This method enables visual readouts without instruments, making it potentially applicable to primary healthcare settings and field surveillance.
Keywords: CRISPR/Cas13a; antibiotic resistance; lateral flow strip; mcr-1; polymyxin.
Copyright © 2025 Song, Hu, Han, Liu, Huang, Niu, Dong, Yan, Jin, Li and Sun.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

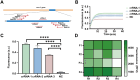


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

