Brucella infection induces chromatin restructuring in host cells to activate immune responses
- PMID: 40539063
- PMCID: PMC12176892
- DOI: 10.3389/fimmu.2025.1574006
Brucella infection induces chromatin restructuring in host cells to activate immune responses
Abstract
Background: Brucella spp., facultative intracellular pathogens that cause brucellosis, drive pathogenesis by invading host cells and establishing intracellular persistence. While their molecular mechanisms are well-characterized, how Brucella induces chromatin restructuring in host cells remains poorly understood, representing a critical gap in host-pathogen interaction research.
Methods: Using an established in vitro infection model of Brucella-infected RAW264.7 murine macrophages, we integrated Hi-C, ATAC-seq, and RNA-seq to generate multi-omics datasets. Multidimensional comparative genomics approaches were employed to systematically map infection-induced changes in host chromatin architecture and functional genomic organization.
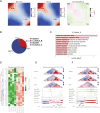
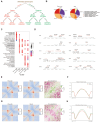
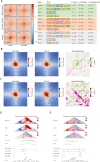
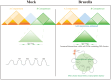
Results: Our findings unveiled substantial alterations in the host chromatin architecture, characterized by a reduction in B-B compartment regions interactions, an increase in A-B compartment interactions, and diminished long-range chromatin contacts. Crucially, Brucella reshaped chromatin compartmentalization, activating interferon-stimulated genes (ISGs) in regions transitioning from compartment B to A. Enhanced sub-TADs interactions within ISG clusters further facilitated their coordinated expression. Additionally, infection remodeled chromatin loop structures, strengthening interactions linked to immune-related gene activation.
Conclusion: These results demonstrate that host cells undergo substantial chromatin remodeling during acute Brucella infection as a defense mechanism against pathogen invasion. Our findings provide critical insights into host-pathogen interactions and suggest potential epigenetic targets for managing brucellosis.
Keywords: 3D genome; Brucella; chromatin restructuring; host-pathogen interactions; interferon-stimulated genes.
Copyright © 2025 Xie, Xu, Su, Lu, Shen, Li, Ye, Hou, Deng, Zhang, Li and Zhao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

